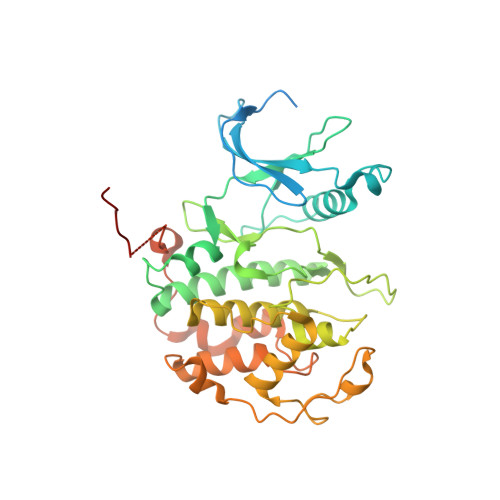
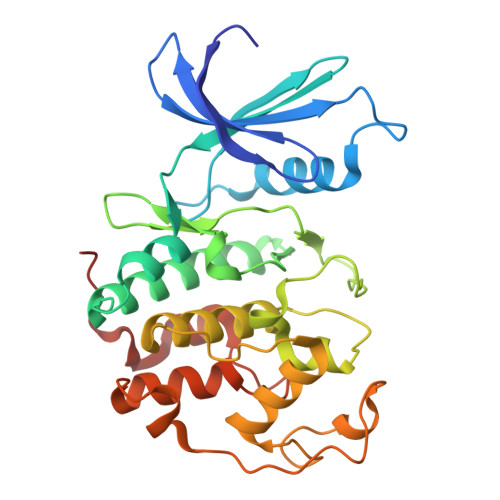
Structural basis of T-loop-independent recognition and activation of CDKs by the CDK-activating kinase.
Cushing, V.I., McGeoch, A.J.S., Williams, S.L., Roumeliotis, T.I., Feng, J., Dan, L.M., Choudhary, J.S., Davey, N.E., Greber, B.J.(2025) Science 390: 911-917
- PubMed: 41100585 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.adw0053
- Primary Citation Related Structures:
9I9I, 9I9J, 9I9K, 9QCV, 9QCX, 9SKQ - PubMed Abstract:
Cyclin-dependent kinases (CDKs) are prototypical regulators of the cell cycle. The CDK-activating kinase (CAK) acts as a master regulator of CDK activity by catalyzing the activating phosphorylation of CDKs on a conserved threonine residue within the regulatory T-loop. However, structural data illuminating the mechanism by which the CAK recognizes and activates CDKs have remained elusive. Here, we determine high-resolution structures of the CAK in complex with CDK2 and CDK2-cyclin A2 by cryogenic electron microscopy. Our structures reveal a T-loop-independent kinase-kinase interface with contributions from both kinase lobes. Computational analysis and structures of CAK in complex with CDK1-cyclin B1 and CDK11 indicate that these structures represent the general architecture of CAK-CDK complexes. These results advance our mechanistic understanding of cell cycle regulation and kinase signaling cascades.
- Division of Structural Biology, The Institute of Cancer Research, 237 Fulham Road, London, UK.
Organizational Affiliation: