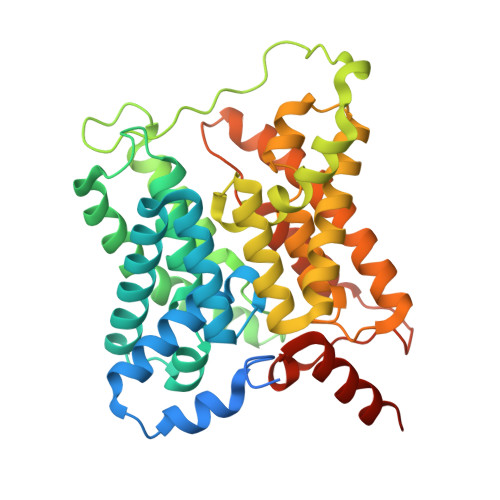
Structural basis of the bifunctionality of Marinobacter salinexigens ZYF650 T glucosylglycerol phosphorylase in glucosylglycerol catabolism.
Lu, D., Zhang, K., Cheng, C., Wu, D., Yin, L., Luo, Q., Shi, M., Ma, H., Lu, X.(2025) J Biological Chem 301: 108127-108127
- PubMed: 39725037 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2024.108127
- Primary Citation Related Structures:
9J1U, 9J22, 9J24, 9J25 - PubMed Abstract:
2-O-α-Glucosylglycerol (GG) is a natural heteroside synthesized by many cyanobacteria and a few heterotrophic bacteria under salt stress conditions. Bacteria produce GG in response to stimuli and degrade it once the stimulus diminishes. Heterotrophic bacteria utilize GG phosphorylase (GGP), a member of the GH13_18 family, via a two-step process consisting of phosphorolysis and hydrolysis for GG catabolism. However, the precise mechanism by which GGP degrades GG remains elusive. We determined the 3D structure of a recently identified GGP (MsGGP) of the deep-sea bacterium Marinobacter salinexigens ZYF650 T , in complex with glucose and glycerol, α-d-glucose-1-phosphate (αGlc1-P), and orthophosphate (inorganic phosphate) at resolutions of 2.5, 2.7, and 2.7 Å, respectively. Notably, the first αGlc1-P complex structure in the GH13_18 family, the complex of MsGGP and αGlc1-P, validates that GGP catalyzes GG decomposition through consecutive phosphorolysis and hydrolysis. In addition, the structure reveals the mechanism of high stereoselectivity on αGlc1-P. Glu231 and Asp190 were identified as the catalytic residues. Interestingly, these structures closely resemble each other, indicating minimal conformational changes upon binding end-product glucose and glycerol, or the intermediate αGlc1-P. The structures also indicate that the substrates may follow a specific trajectory and a precise order toward the active center in close proximity and in a geometrically favorable orientation for catalysis in a double displacement mechanism.
- School of Chemical Engineering, Marine and Life Sciences, Dalian University of Technology, Panjin, China; Key Laboratory of Biofuels, Qingdao Institute of Bioenergy and Bioprocess Technology, Chinese Academy of Sciences, Qingdao, China.
Organizational Affiliation: