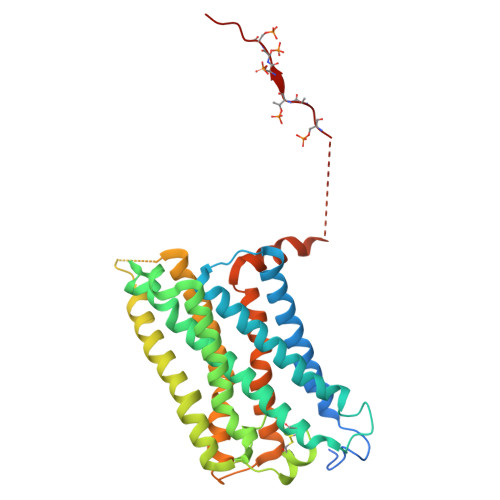
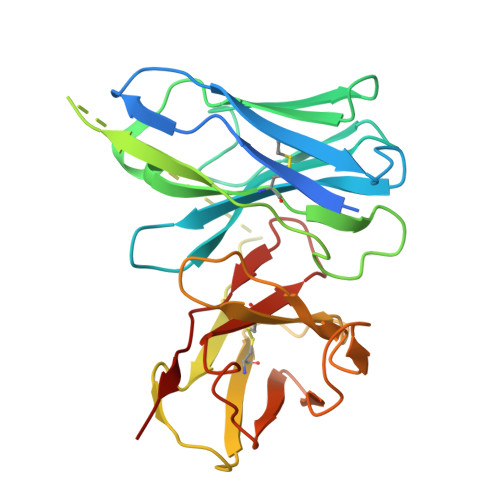
Constitutive arrestin recruitment by orphan GPR52 via an atypical binding mode.
Lin, X., Wei, X., Pu, N., Wang, L., Zhang, Z., Li, C., Yue, Y., Liu, J., Tan, Q., Sun, Q., Xu, F.(2025) Cell Res 35: 913-916
- PubMed: 40813798 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-025-01165-w
- Primary Citation Related Structures:
9IJR, 9IJS - iHuman Institute, ShanghaiTech University, Shanghai, China. linxi@shanghaitech.edu.cn.
Organizational Affiliation: