High-throughput investigation of cyclin docking interactions reveals the complexity of motif binding determinants.
Ord, M., Winters, M.J., Subbanna, M.S., de Martin Garrido, N., Cushing, V.I., Kliche, J., Benz, C., Ivarsson, Y., Greber, B.J., Pryciak, P.M., Davey, N.E.(2025) Nat Commun 16: 7622-7622
- PubMed: 40817109
- DOI: https://doi.org/10.1038/s41467-025-62765-z
- Primary Citation Related Structures:
9HIU, 9HIW, 9HJ1 - PubMed Abstract:
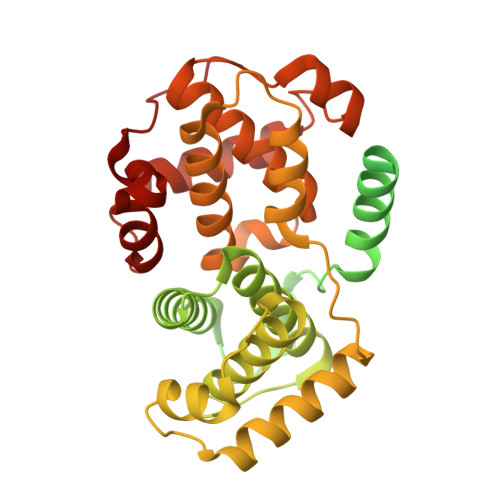
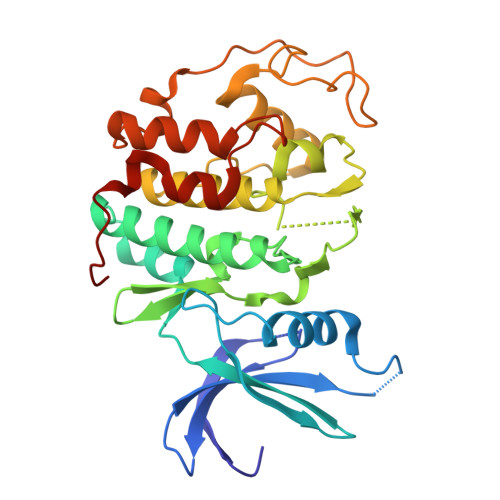
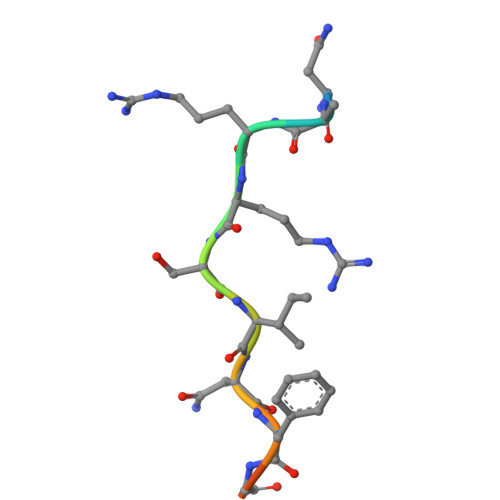
Many regulatory protein-protein interactions depend on Short Linear Motifs (SLiMs). In the cell cycle, cyclin-CDKs recognize SLiMs to control substrate recruitment and phosphorylation timing. Here, we measure the relative binding strength of ~100,000 peptides to 11 human cyclins from five families (D, E, A, B, and F). Using a quantitative intracellular binding assay and large-scale tiled peptide screening, we identify multiple non-canonical binders unveiling a broader repertoire of cyclin docking motif types. Cryo-electron microscopy and saturation mutagenesis studies reveal distinct binding modes and sequence features governing motif recognition, binding strength, and cyclin preference. Docking motifs vary from highly selective to pan-cyclin, thereby fine-tuning the timing of CDK phosphorylation during cell cycle. Overall, these findings provide insights into the rules encoding specificity and affinity of SLiM-mediated interactions and offer a framework for understanding motif-driven protein networks across the proteome.
- University of Cambridge, CRUK Cambridge Institute, Cambridge, UK.
Organizational Affiliation: