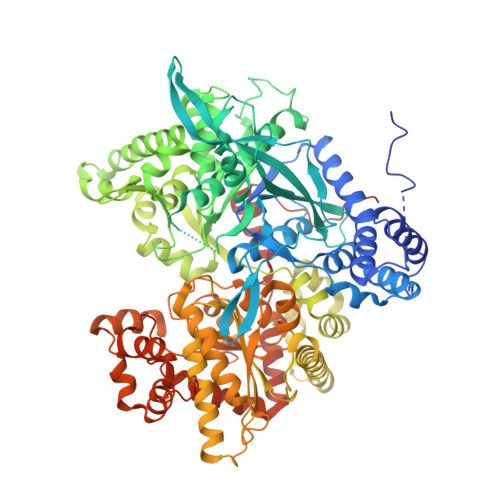
The allosteric transition of glycogen phosphorylase.
Barford, D., Johnson, L.N.(1989) Nature 340: 609-616
- PubMed: 2770867 Search on PubMed
- DOI: https://doi.org/10.1038/340609a0
- Primary Citation Related Structures:
9GPB - PubMed Abstract:
The crystal structure of R-state glycogen phosphorylase b has been determined at 2.9 A resolution. A comparison of T-state and R-state structures of the enzyme explains its cooperative behaviour on ligand binding and the allosteric regulation of its activity. Communication between catalytic sites of the dimer is provided by a change in packing geometry of two helices linking each site with the subunit interface. Activation by AMP or by phosphorylation results in a quaternary conformational change that switches these two helices into the R-state conformation.
- Laboratory of Molecular Biophysics, University of Oxford, UK.
Organizational Affiliation: