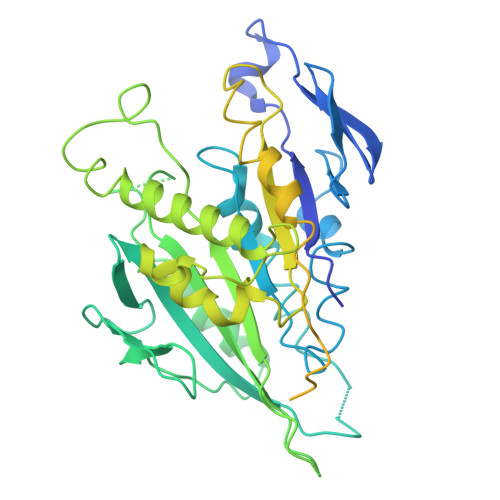
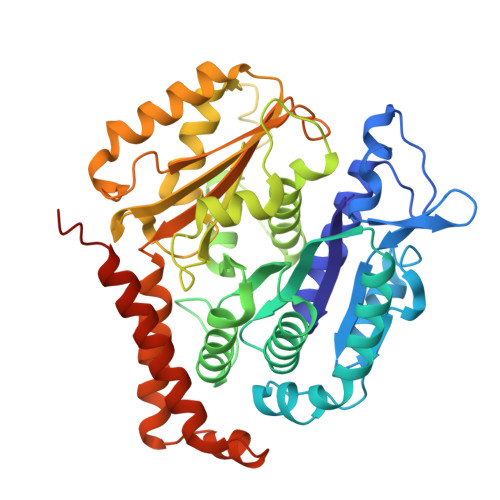
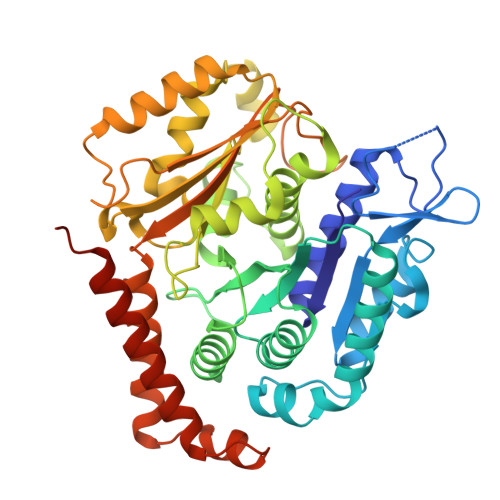
HURP regulates Kif18A recruitment and activity to synergistically control microtubule dynamics.
Perez-Bertoldi, J.M., Zhao, Y., Thawani, A., Yildiz, A., Nogales, E.(2024) Nat Commun 15: 9687-9687
- PubMed: 39516196 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-53691-7
- Primary Citation Related Structures:
9DHZ, 9DI0, 9DXC, 9DXE - PubMed Abstract:
During mitosis, microtubule dynamics are regulated to ensure proper alignment and segregation of chromosomes. The dynamics of kinetochore-attached microtubules are regulated by hepatoma-upregulated protein (HURP) and the mitotic kinesin-8 Kif18A, but the underlying mechanism remains elusive. Using single-molecule imaging in vitro, we demonstrate that Kif18A motility is regulated by HURP. While sparse decoration of HURP activates the motor, higher concentrations hinder processive motility. To shed light on this behavior, we determine the binding mode of HURP to microtubules using cryo-EM. The structure helps rationalize why HURP functions as a microtubule stabilizer. Additionally, HURP partially overlaps with the microtubule-binding site of the Kif18A motor domain, indicating that excess HURP inhibits Kif18A motility by steric exclusion. We also observe that HURP and Kif18A function together to suppress dynamics of the microtubule plus-end, providing a mechanistic basis for how they collectively serve in microtubule length control.
- Biophysics Graduate Group, University of California, Berkeley, CA, USA.
Organizational Affiliation: