Structural divergence of H2A.Z-associated human chromatin remodelers SRCAP and TIP60
Park, G., Patel, A.B., Wu, C., Louder, R.K.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| E1A-binding protein p400,E1A-binding protein p400/Haloalkane dehalogenase chimera | 2,841 | Homo sapiens, Rhodococcus rhodochrous This entity is chimeric | Mutation(s): 0 Gene Names: EP400, CAGH32, KIAA1498, KIAA1818, TNRC12, dhaA EC: 3.6.4 (PDB Primary Data), 3.8.1.5 (PDB Primary Data) | 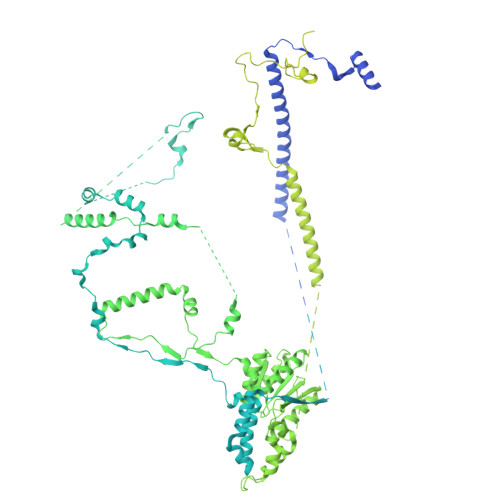 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q96L91 GTEx: ENSG00000183495 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q96L91 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Vacuolar protein sorting-associated protein 72 homolog | 364 | Homo sapiens | Mutation(s): 0 Gene Names: VPS72, TCFL1, YL1 | 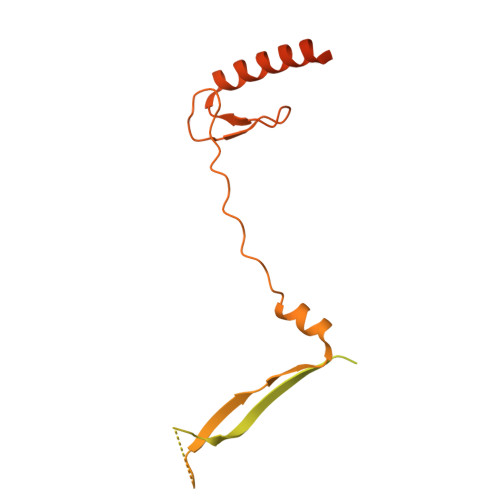 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q15906 GTEx: ENSG00000163159 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q15906 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Enhancer of polycomb homolog 1 | 263 | Homo sapiens | Mutation(s): 0 Gene Names: EPC1 |  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9H2F5 GTEx: ENSG00000120616 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9H2F5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| RuvB-like 1 | D [auth E], F [auth G], H [auth I] | 456 | Homo sapiens | Mutation(s): 0 Gene Names: RUVBL1, INO80H, NMP238, TIP49, TIP49A EC: 3.6.4.12 | 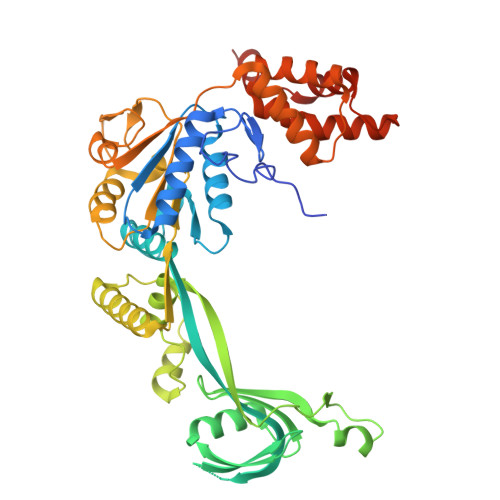 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9Y265 GTEx: ENSG00000175792 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9Y265 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| RuvB-like 2,RuvB-like 2/Maltose/maltodextrin-binding periplasmic protein chimera | E [auth F], G [auth H], I [auth J] | 857 | Homo sapiens, Escherichia coli K-12 This entity is chimeric | Mutation(s): 0 Gene Names: RUVBL2, INO80J, TIP48, TIP49B, CGI-46, malE, b4034, JW3994 EC: 3.6.4.12 | 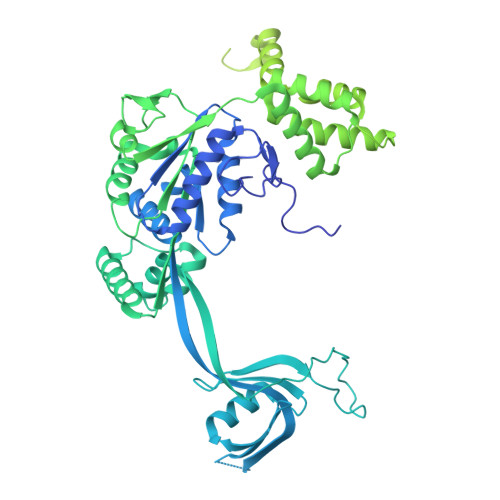 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9Y230 GTEx: ENSG00000183207 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9Y230 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 6 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Actin, cytoplasmic 1 | J [auth L] | 375 | Homo sapiens | Mutation(s): 0 Gene Names: ACTB EC: 3.6.4 |  |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P60709 GTEx: ENSG00000075624 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P60709 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 7 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Actin-like protein 6A | K [auth M] | 429 | Homo sapiens | Mutation(s): 0 Gene Names: ACTL6A, BAF53, BAF53A, INO80K |  |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O96019 GTEx: ENSG00000136518 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O96019 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 8 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA methyltransferase 1-associated protein 1 | L [auth N] | 467 | Homo sapiens | Mutation(s): 0 Gene Names: DMAP1, KIAA1425 |  |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9NPF5 GTEx: ENSG00000178028 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9NPF5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| AGS Download:Ideal Coordinates CCD File | AA [auth M] M [auth E] O [auth F] Q [auth G] S [auth H] | PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER C10 H16 N5 O12 P3 S NLTUCYMLOPLUHL-KQYNXXCUSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | BA [auth M] N [auth E] P [auth F] R [auth G] T [auth H] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.21_5207 |
| RECONSTRUCTION | RELION |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | -- |