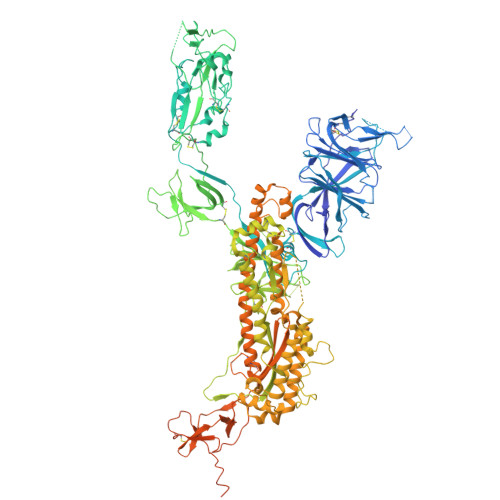
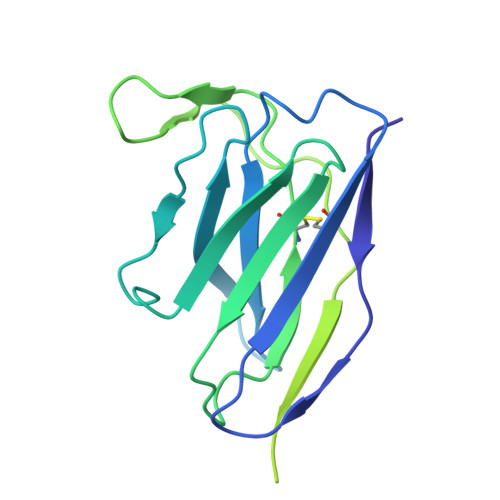
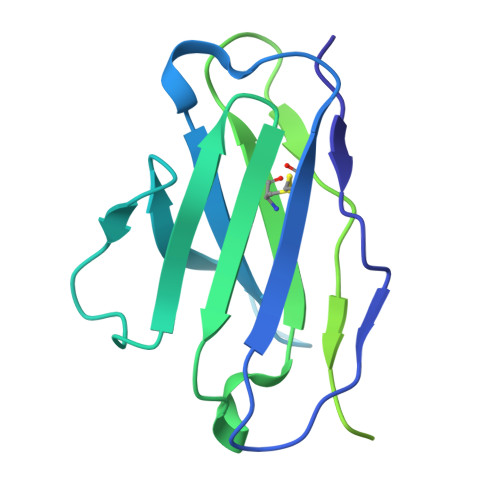
Bispecific antibodies targeting the N-terminal and receptor binding domains potently neutralize SARS-CoV-2 variants of concern.
Rubio, A.A., Baharani, V.A., Dadonaite, B., Parada, M., Abernathy, M.E., Wang, Z., Lee, Y.E., Eso, M.R., Phung, J., Ramos, I., Chen, T., El Nesr, G., Bloom, J.D., Bieniasz, P.D., Nussenzweig, M.C., Barnes, C.O.(2025) Sci Transl Med 17: eadq5720-eadq5720
- PubMed: 40043139 Search on PubMed
- DOI: https://doi.org/10.1126/scitranslmed.adq5720
- Primary Citation Related Structures:
9BJ2, 9BJ3, 9BJ4 - PubMed Abstract:
The ongoing emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants of concern (VOCs) that reduce the effectiveness of antibody therapeutics necessitates development of next-generation antibody modalities that are resilient to viral evolution. Here, we characterized amino-terminal domain (NTD)- and receptor binding domain (RBD)-specific monoclonal antibodies previously isolated from coronavirus disease 2019 (COVID-19) convalescent donors for their activity against emergent SARS-CoV-2 VOCs. Among these, the NTD-specific antibody C1596 displayed the greatest breadth of binding to VOCs, with cryo-electron microscopy structural analysis revealing recognition of a distinct NTD epitope outside of the site i antigenic supersite. Given C1596's favorable binding profile, we designed a series of bispecific antibodies (bsAbs), termed CoV2-biRNs, that featured both NTD and RBD specificities. Two of the C1596-inclusive bsAbs, CoV2-biRN5 and CoV2-biRN7, retained potent in vitro neutralization activity against all Omicron variants tested, including XBB.1.5, BA.2.86, and JN.1, contrasting the diminished potency of parental antibodies delivered as monotherapies or as a cocktail. Furthermore, prophylactic delivery of CoV2-biRN5 reduced the viral load within the lungs of K18-hACE2 mice after challenge with SARS-CoV-2 XBB.1.5. In conclusion, NTD-RBD bsAbs offer promising potential for the design of resilient, next-generation antibody therapeutics against SARS-CoV-2 VOCs.
- Stanford Biosciences, Stanford School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: