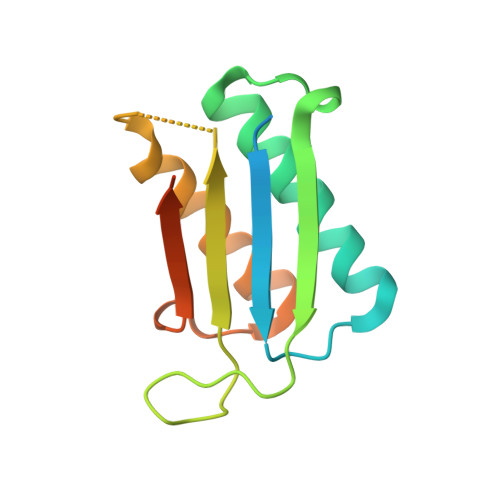
Crystal structure of Plasmodium vivax macrophage migration inhibitory factor
Nair, A., Lin, M., Srivastava, A., Liu, L., Cooper, A., Battaile, K.P., Harmon, E., Myler, P.J., Staker, B.L., Lovell, S., Chakafana, G., Asojo, O.A.(2026) Acta Crystallogr F Struct Biol Commun