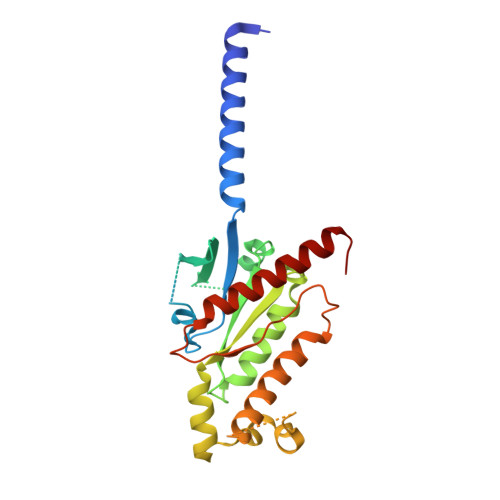
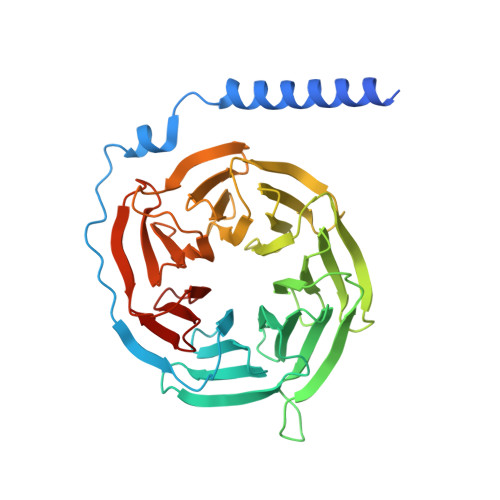
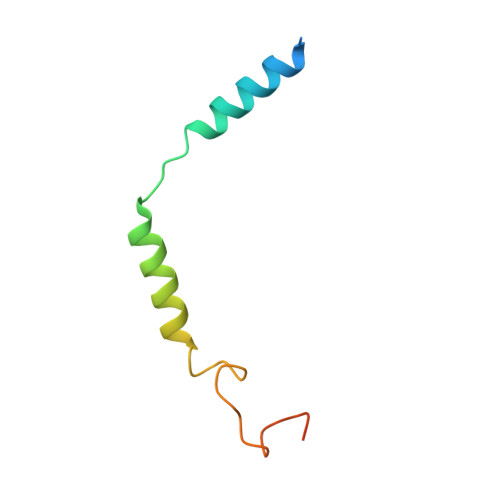
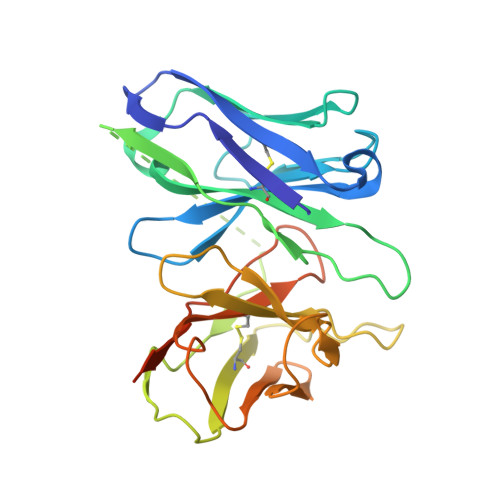
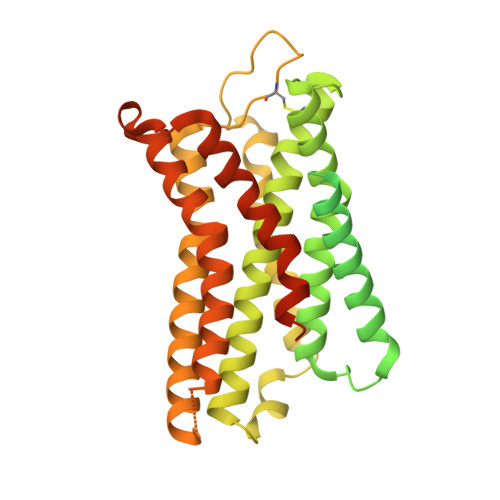
Structural basis of agonist specificity of alpha 1A -adrenergic receptor.
Su, M., Wang, J., Xiang, G., Do, H.N., Levitz, J., Miao, Y., Huang, X.Y.(2023) Nat Commun 14: 4819-4819
- PubMed: 37563160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-40524-2
- Primary Citation Related Structures:
8THK, 8THL - PubMed Abstract:
α 1 -adrenergic receptors (α 1 -ARs) play critical roles in the cardiovascular and nervous systems where they regulate blood pressure, cognition, and metabolism. However, the lack of specific agonists for all α 1 subtypes has limited our understanding of the physiological roles of different α 1 -AR subtypes, and led to the stagnancy in agonist-based drug development for these receptors. Here we report cryo-EM structures of α 1A -AR in complex with heterotrimeric G-proteins and either the endogenous common agonist epinephrine or the α 1A -AR-specific synthetic agonist A61603. These structures provide molecular insights into the mechanisms underlying the discrimination between α 1A -AR and α 1B -AR by A61603. Guided by the structures and corresponding molecular dynamics simulations, we engineer α 1A -AR mutants that are not responsive to A61603, and α 1B -AR mutants that can be potently activated by A61603. Together, these findings advance our understanding of the agonist specificity for α 1 -ARs at the molecular level, opening the possibility of rational design of subtype-specific agonists.
- Department of Physiology and Biophysics, Weill Cornell Medical College of Cornell University, New York, NY, 10065, USA.
Organizational Affiliation: