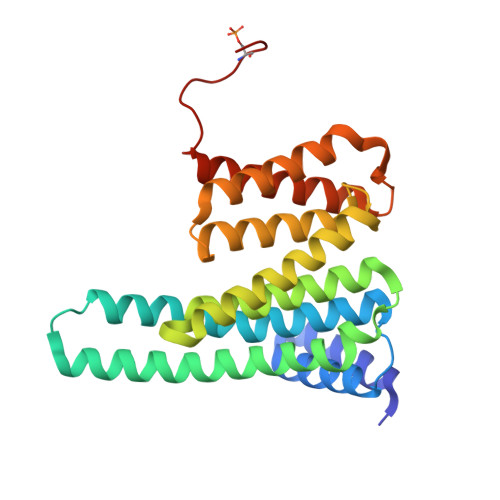
Crystal structure reveals canonical recognition of the phosphorylated cytoplasmic loop of human alpha7 nicotinic acetylcholine receptor by 14-3-3 protein.
Sluchanko, N.N., Kapitonova, A.A., Shulepko, M.A., Kukushkin, I.D., Kulbatskii, D.S., Tugaeva, K.V., Varfolomeeva, L.A., Minyaev, M.E., Boyko, K.M., Popov, V.O., Kirpichnikov, M.P., Lyukmanova, E.N.(2023) Biochem Biophys Res Commun 682: 91-96
- PubMed: 37804592
- DOI: https://doi.org/10.1016/j.bbrc.2023.09.086
- Primary Citation of Related Structures:
8P1H - PubMed Abstract:
Nicotinic acetylcholine receptors (nAChRs) are ligand-gated ion channels composed of five homologous subunits. The homopentameric α7-nAChR, abundantly expressed in the brain, is involved in the regulation of the neuronal plasticity and memory and undergoes phosphorylation by protein kinase A (PKA). Here, we extracted native α7-nAChR from murine brain, validated its assembly by cryo-EM and showed that phosphorylation by PKA in vitro enables its interaction with the abundant human brain protein 14-3-3ζ. Bioinformatic analysis narrowed the putative 14-3-3-binding site down to the fragment of the intracellular loop (ICL) containing Ser365 (Q 361 RRCSLASVEMS 372 ), known to be phosphorylated in vivo. We reconstructed the 14-3-3ζ/ICL peptide complex and determined its structure by X-ray crystallography, which confirmed the Ser365 phosphorylation-dependent canonical recognition of the ICL by 14-3-3. A common mechanism of nAChRs' regulation by ICL phosphorylation and 14-3-3 binding that potentially affects nAChR activity, stoichiometry, and surface expression is suggested.
- A.N. Bach Institute of Biochemistry, Federal Research Center of Biotechnology of the Russian Academy of Sciences, Moscow, 119071, Russia. Electronic address: nikolai.sluchanko@mail.ru.
Organizational Affiliation: