A potent and broad-spectrum neutralizing nanobody for SARS-CoV-2 viruses, including all major Omicron strains.
Yao, H., Wang, H., Zhang, Z., Lu, Y., Zhang, Z., Zhang, Y., Xiong, X., Wang, Y., Wang, Z., Yang, H., Zhao, J., Xu, W.(2023) MedComm (2020) 4: e397-e397
- PubMed: 37901798 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/mco2.397
- Primary Citation Related Structures:
8K3K, 8K45, 8K46, 8K47 - PubMed Abstract:
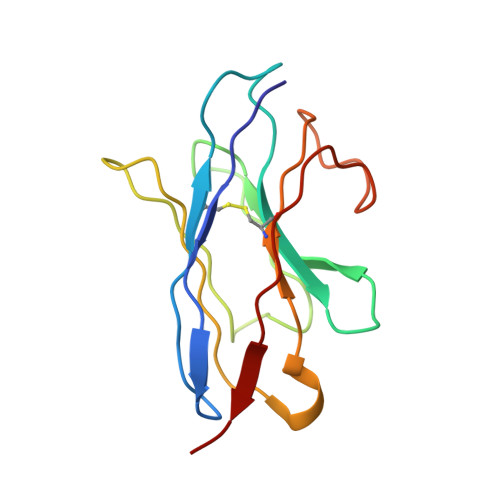
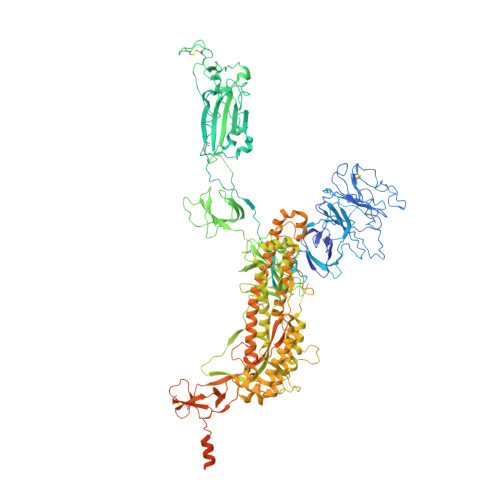
SARS-CoV-2 viruses are highly transmissible and immune evasive. It is critical to develop broad-spectrum prophylactic and therapeutic antibodies for potential future pandemics. Here, we used the phage display method to discover nanobodies (Nbs) for neutralizing SARS-CoV-2 viruses especially Omicron strains. The leading nanobody (Nb), namely, Nb4, with excellent physicochemical properties, can neutralize Delta and Omicron subtypes, including BA.1, BA.1.1 (BA.1 + R346K), BA.2, BA.5, BQ.1, and XBB.1. The crystal structure of Nb4 in complex with the receptor-binding domain (RBD) of BA.1 Spike protein reveals that Nb4 interacts with an epitope on the RBD overlapping with the receptor-binding motif, and thus competes with angiotensin-converting enzyme 2 (ACE2) binding. Nb4 is expected to be effective for neutralizing most recent Omicron variants, since the epitopes are evolutionarily conserved among them. Indeed, trivalent Nb4 interacts with the XBB1.5 Spike protein with low nM affinity and competes for ACE2 binding. Prophylactic and therapeutic experiments in mice indicated that Nb4 could reduce the Omicron virus loads in the lung. In particular, in prophylactic experiments, intranasal administration of multivalent Nb4 completely protected mice from Omicron infection. Taken together, these results demonstrated that Nb4 could serve as a potent and broad-spectrum prophylactic and therapeutic Nb for COVID-19.
- School of Life Science and Technology ShanghaiTech University Shanghai China.
Organizational Affiliation: