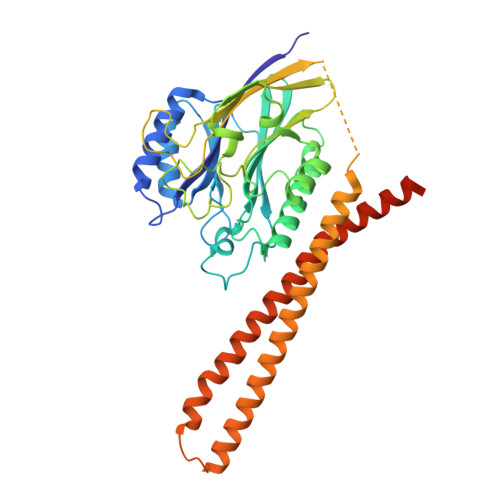
Molecular basis for the catalytic mechanism of human neutral sphingomyelinases 1 (hSMPD2).
Yi, J., Qi, B., Yin, J., Li, R., Chen, X., Hu, J., Li, G., Zhang, S., Zhang, Y., Yang, M.(2023) Nat Commun 14: 7755-7755
- PubMed: 38012235 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-43580-w
- Primary Citation Related Structures:
8J2F - PubMed Abstract:
Enzymatic breakdown of sphingomyelin by sphingomyelinase (SMase) is the main source of the membrane lipids, ceramides, which are involved in many cellular physiological processes. However, the full-length structure of human neutral SMase has not been resolved; therefore, its catalytic mechanism remains unknown. Here, we resolve the structure of human full-length neutral SMase, sphingomyelinase 1 (SMPD2), which reveals that C-terminal transmembrane helices contribute to dimeric architecture of hSMPD2 and that D111 - K116 loop domain is essential for substrate hydrolysis. Coupled with molecular docking, we clarify the binding pose of sphingomyelin, and site-directed mutagenesis further confirms key residues responsible for sphingomyelin binding. Hybrid quantum mechanics/molecular mechanics (QM/MM) molecular dynamic (MD) simulations are utilized to elaborate the catalysis of hSMPD2 with the reported in vitro substrates, sphingomyelin and lyso-platelet activating fator (lyso-PAF). Our study provides mechanistic details that enhance our knowledge of lipid metabolism and may lead to an improved understanding of ceramide in disease and in cancer treatment.
- Ministry of Education Key Laboratory of Protein Science, Tsinghua-Peking Joint Center for Life Sciences, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing, 100084, China.
Organizational Affiliation: