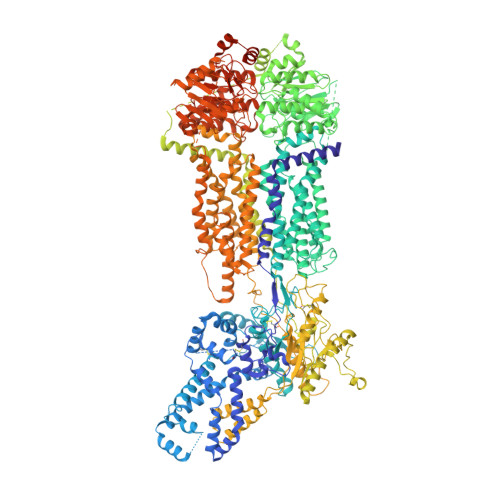
Cryo-EM structures of human ABCA7 provide insights into its phospholipid translocation mechanisms.
Le, L.T.M., Thompson, J.R., Dehghani-Ghahnaviyeh, S., Pant, S., Dang, P.X., French, J.B., Kanikeyo, T., Tajkhorshid, E., Alam, A.(2023) EMBO J 42: e111065-e111065
- PubMed: 36484366 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2022111065
- Primary Citation Related Structures:
8EDW, 8EE6, 8EEB, 8EOP - PubMed Abstract:
Phospholipid extrusion by ABC subfamily A (ABCA) exporters is central to cellular physiology, although the specifics of the underlying substrate interactions and transport mechanisms remain poorly resolved at the molecular level. Here we report cryo-EM structures of lipid-embedded human ABCA7 in an open state and in a nucleotide-bound, closed state at resolutions between 3.6 and 4.0 Å. The former reveals an ordered patch of bilayer lipids traversing the transmembrane domain (TMD), while the latter reveals a lipid-free, closed TMD with a small extracellular opening. These structures offer a structural framework for both substrate entry and exit from the ABCA7 TMD and highlight conserved rigid-body motions that underlie the associated conformational transitions. Combined with functional analysis and molecular dynamics (MD) simulations, our data also shed light on lipid partitioning into the ABCA7 TMD and localized membrane perturbations that underlie ABCA7 function and have broader implications for other ABCA family transporters.
- The Hormel Institute, University of Minnesota, Austin, MN, USA.
Organizational Affiliation: