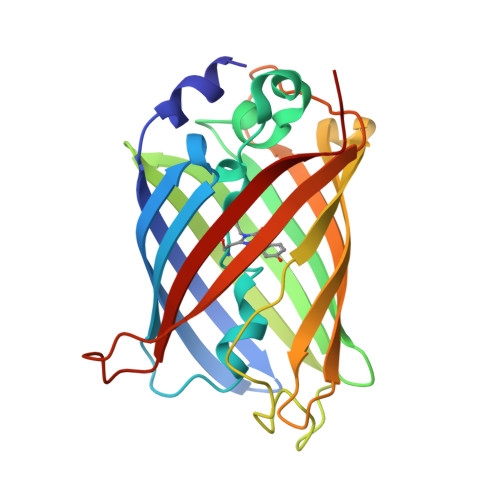
Structural characterization of green fluorescent protein in the I-state.
Takeda, R., Tsutsumi, E., Okatsu, K., Fukai, S., Takeda, K.(2024) Sci Rep 14: 22832-22832
- PubMed: 39353998 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-024-73696-y
- Primary Citation Related Structures:
8ZUP, 8ZUQ, 8ZUR, 8ZUS, 8ZUT - PubMed Abstract:
Green fluorescent protein (GFP) is widely utilized as a fluorescent tag in biochemical fields. Whereas the intermediate (I) state has been proposed in the photoreaction cycle in addition to the A and B states, until now the structure of I has only been estimated by computational studies. In this paper, we report the crystal structures of the I stabilizing variants of GFP at high resolutions where respective atoms can be observed separately. Comparison with the structures in the other states highlights the structural feature of the I state. The side chain of one of the substituted residues, Val203, adopts the gauche- conformation observed for Thr203 in the A state, which is different from the B state. On the other hand, His148 interacts with the chromophore by ordinary hydrogen bonding with a distance of 2.85 Å, while the weaker interaction by longer distances is observed in the A state. Therefore, it was indicated that it is possible to distinguish three states A, B and I by the two hydrogen bond distances Oγ-Thr203···Oη-chromophore and Nδ1-His148···Oη-chromophore. We discuss the characteristics of the I intermediate of wild-type GFP on the bases of the structure estimated from the variant structures by quantum chemical calculations.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto, 606-8502, Japan.
Organizational Affiliation: