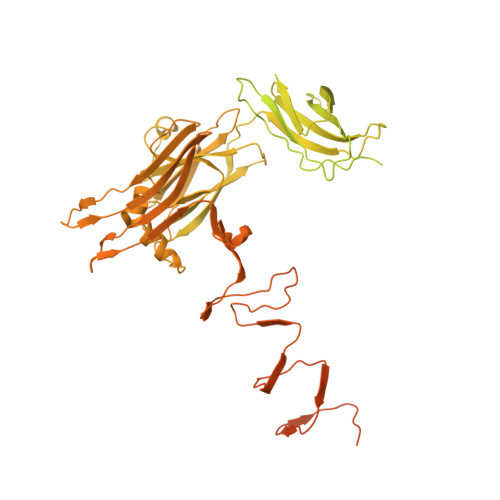
Structural mechanism of bacteriophage lambda tail's interaction with the bacterial receptor.
Ge, X., Wang, J.(2024) Nat Commun 15: 4185-4185
- PubMed: 38760367 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-48686-3
- Primary Citation Related Structures:
8XCG, 8XCI, 8XCJ, 8XCK - PubMed Abstract:
Bacteriophage infection, a pivotal process in microbiology, initiates with the phage's tail recognizing and binding to the bacterial cell surface, which then mediates the injection of viral DNA. Although comprehensive studies on the interaction between bacteriophage lambda and its outer membrane receptor, LamB, have provided rich information about the system's biochemical properties, the precise molecular mechanism remains undetermined. This study revealed the high-resolution cryo-electron microscopy (cryo-EM) structures of the bacteriophage lambda tail complexed with its irreversible Shigella sonnei 3070 LamB receptor and the closed central tail fiber. These structures reveal the complex processes that trigger infection and demonstrate a substantial conformational change in the phage lambda tail tip upon LamB binding. Providing detailed structures of bacteriophage lambda infection initiation, this study contributes to the expanding knowledge of lambda-bacterial interaction, which holds significance in the fields of microbiology and therapeutic development.
- State Key Laboratory of Membrane Biology, Beijing Frontier Research Center for Biological Structure, School of Life Sciences, Tsinghua University, 100084, Beijing, PR China.
Organizational Affiliation: