A Compact Reprogrammed Genetic Code for De Novo Discovery of Proteolytically Stable Thiopeptides.
Vinogradov, A.A., Zhang, Y., Hamada, K., Kobayashi, S., Ogata, K., Sengoku, T., Goto, Y., Suga, H.(2024) J Am Chem Soc 146: 8058-8070
- PubMed: 38491946 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.3c12037
- Primary Citation Related Structures:
8WM0 - PubMed Abstract:
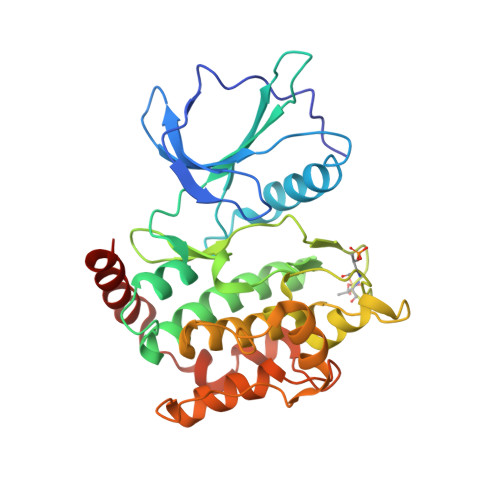
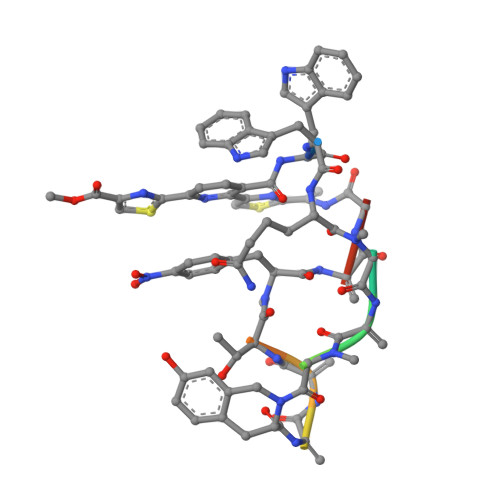
Thiopeptides make up a group of structurally complex peptidic natural products holding promise in bioengineering applications. The previously established thiopeptide/mRNA display platform enables de novo discovery of natural product-like thiopeptides with designed bioactivities. However, in contrast to natural thiopeptides, the discovered structures are composed predominantly of proteinogenic amino acids, which results in low metabolic stability in many cases. Here, we redevelop the platform and demonstrate that the utilization of compact reprogrammed genetic codes in mRNA display libraries can lead to the discovery of thiopeptides predominantly composed of nonproteinogenic structural elements. We demonstrate the feasibility of our designs by conducting affinity selections against Traf2- and NCK-interacting kinase (TNIK). The experiment identified a series of thiopeptides with high affinity to the target protein (the best K D = 2.1 nM) and kinase inhibitory activity (the best IC 50 = 0.15 μM). The discovered compounds, which bore as many as 15 nonproteinogenic amino acids in an 18-residue macrocycle, demonstrated high metabolic stability in human serum with a half-life of up to 99 h. An X-ray cocrystal structure of TNIK in complex with a discovered thiopeptide revealed how nonproteinogenic building blocks facilitate the target engagement and orchestrate the folding of the thiopeptide into a noncanonical conformation. Altogether, the established platform takes a step toward the discovery of thiopeptides with high metabolic stability for early drug discovery applications.
- Department of Chemistry, Graduate School of Science, The University of Tokyo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: