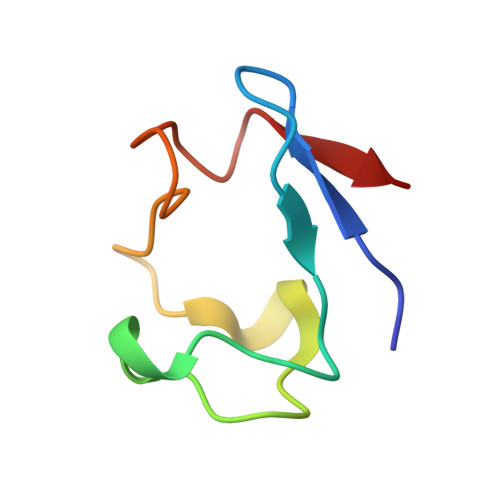
Refinement of rubredoxin from Desulfovibrio vulgaris at 1.0 A with and without restraints.
Dauter, Z., Sieker, L.C., Wilson, K.S.(1992) Acta Crystallogr B 48: 42-59
- PubMed: 1616692
- DOI: https://doi.org/10.1107/s0108768191010613
- Primary Citation Related Structures:
8RXN - PubMed Abstract:
X-ray data have been recorded from crystals of rubredoxin derived from the bacterium Desulfovibrio vulgaris to a resolution of 1.0 A using in part synchrotron radiation and in part X-rays from a sealed-tube Mo K alpha source. In both cases an imaging-plate scanner was used as detector. The space group of the crystals is P2(1) with cell dimensions a = 19.97, b = 41.45, c = 24.41 A and beta = 108.3 degrees. The overall merging R(I) factor between symmetry-related reflections was 5.8%. The model was refined by least-squares minimization initially with stereochemical restraints to an R factor of 16.4%. Only atomic positional parameters and isotropic temperature factors for non-H atoms were used in the refinement. There were 18,532 independent X-ray observations for a total of 1916 atomic parameters. A round of unrestrained refinement gave an R factor of 16.0%, acceptable geometry for more than 90% of the protein atoms, but emphasized the disorder inherent in eight of the residues. A final round of restrained refinement gave an R factor of 14.7%. Three of the 389 protein atoms in the molecule, in the side chain of Lys2, have been assigned zero occupancy in the model. A total of eight atoms in three side chains have been assigned two conformations, giving 393 protein atomic sites in the model. In addition there is one Fe atom, a sulfate ion and 102 water sites. 339 H atoms were included at their calculated positions, which were not refined. There is clear evidence for anisotropic thermal motion. This has not been incorporated in the present model.
- European Molecular Biology Laboratory (EMBL), Hamburg, Germany.
Organizational Affiliation: