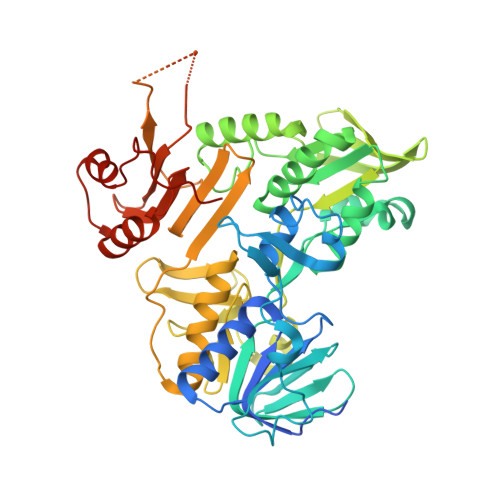
CHCHD4 binding affects the active site of apoptosis inducing factor (AIF): Structural determinants for allosteric regulation.
Fagnani, E., Cocomazzi, P., Pellegrino, S., Tedeschi, G., Scalvini, F.G., Cossu, F., Da Vela, S., Aliverti, A., Mastrangelo, E., Milani, M.(2024) Structure 32: 594-602.e4
- PubMed: 38460521 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2024.02.008
- Primary Citation Related Structures:
8QNS - PubMed Abstract:
Apoptosis-inducing factor (AIF), which is confined to mitochondria of normal healthy cells, is the first identified caspase-independent cell death effector. Moreover, AIF is required for the optimal functioning of the respiratory chain machinery. Recent findings have revealed that AIF fulfills its pro-survival function by interacting with CHCHD4, a soluble mitochondrial protein which promotes the entrance and the oxidative folding of different proteins in the inner membrane space. Here, we report the crystal structure of the ternary complex involving the N-terminal 27-mer peptide of CHCHD4, NAD + , and AIF harboring its FAD (flavin adenine dinucleotide) prosthetic group in oxidized form. Combining this information with biophysical and biochemical data on the CHCHD4/AIF complex, we provide a detailed structural description of the interaction between the two proteins, validated by both chemical cross-linking mass spectrometry analysis and site-directed mutagenesis.
- Biophysics Institute, CNR-IBF, Via Corti 12, 20133 Milan, Italy; Department of Bioscience, Università degli Studi di Milano, Via Celoria 26, 20133 Milan, Italy.
Organizational Affiliation: