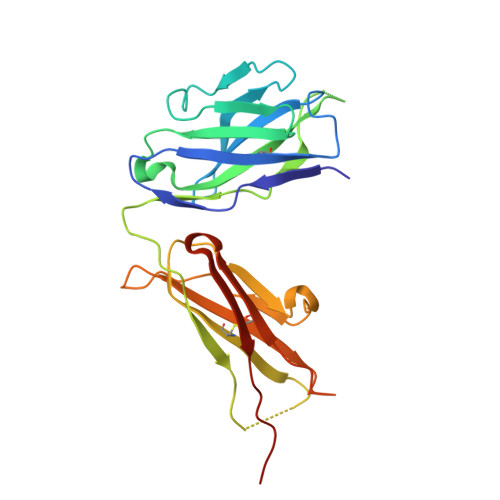
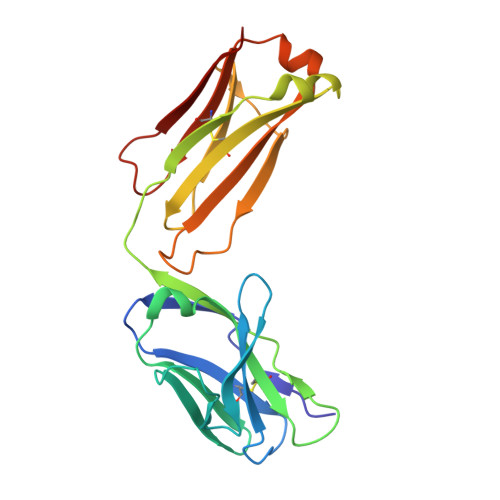
Structure-guided disulfide engineering restricts antibody conformation to elicit TNFR agonism.
Elliott, I.G., Fisher, H., Chan, H.T.C., Inzhelevskaya, T., Mockridge, C.I., Penfold, C.A., Duriez, P.J., Orr, C.M., Herniman, J., Muller, K.T.J., Essex, J.W., Cragg, M.S., Tews, I.(2025) Nat Commun 16: 3495-3495
- PubMed: 40221417 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-58773-8
- Primary Citation Related Structures:
8PUK, 8PUL - PubMed Abstract:
A promising strategy in cancer immunotherapy is activation of immune signalling pathways through antibodies that target co-stimulatory receptors. hIgG2, one of four human antibody isotypes, is known to deliver strong agonistic activity, and modification of hIgG2 hinge disulfides can influence immune-stimulating activity. This was shown for antibodies directed against the hCD40 receptor, where cysteine-to-serine exchange mutations caused changes in antibody conformational flexibility. Here we demonstrate that the principles of increasing agonism by restricting antibody conformation through disulfide modification can be translated to the co-stimulatory receptor h4-1BB, another member of the tumour necrosis factor receptor superfamily. Furthermore, we explore structure-guided design of the anti-hCD40 antibody ChiLob7/4 and show that engineering additional disulfides between opposing F(ab') arms can elicit conformational restriction, concomitant with enhanced agonism. These results support a mode where subtle increases in rigidity can deliver significant improvements in immunostimulatory activity, thus providing a strategy for the rational design of more powerful antibody therapeutics.
- School of Chemistry and Chemical Engineering, Faculty of Engineering and Physical Sciences, University of Southampton, Southampton, SO17 1BJ, UK.
Organizational Affiliation: