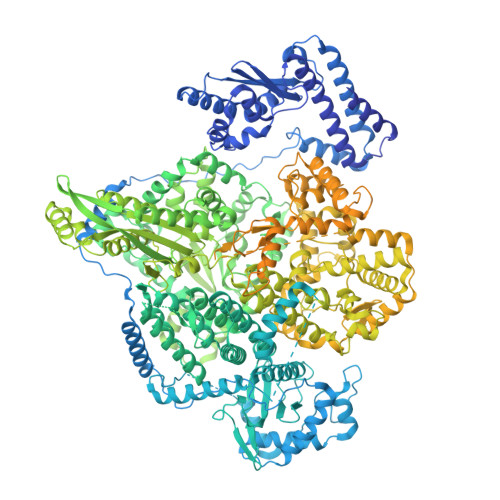
Structural characterization of the full-length Hantaan virus polymerase.
Keown, J.R., Carrique, L., Nilsson-Payant, B.E., Fodor, E., Grimes, J.M.(2024) PLoS Pathog 20: e1012781-e1012781
- PubMed: 39652621
- DOI: https://doi.org/10.1371/journal.ppat.1012781
- Primary Citation Related Structures:
8P1J, 8P1K, 8P1L, 8P1M, 8P1N - PubMed Abstract:
Hantaviridae are a family of segmented negative-sense RNA viruses that contain important human and animal pathogens. Hantaviridae contain a viral RNA-dependent RNA polymerase that replicates and transcribes the viral genome. Here we establish the expression and purification of the polymerase from the Old World Hantaan virus and characterise the structure using Cryo-EM. We determine a series of structures at resolutions between 2.7 and 3.3 Å of RNA free polymerase comprising the core, core and endonuclease, and a full-length polymerase. The full-length polymerase structure depicts the location of the cap binding and C-terminal domains which are arranged in a conformation that is incompatible with transcription and in a novel conformation not observed in previous conformations of cap-snatching viral polymerases. We further describe structures with 5' vRNA promoter in the presence and absence of a nucleotide triphosphate. The nucleotide bound structure mimics a replication pre-initiation complex and the nucleotide stabilises the motif E in a conformation distinct from those previously observed. We observe motif E in four distinct conformations including β-sheet, two helical arrangements, and nucleotide primed arrangement. The insights gained here guide future mechanistic studies of both the transcription and replication activities of the hantavirus polymerase and for the development of therapeutic targets.
- Division of Structural Biology, Centre for Human Genetics, University of Oxford, Oxford, United Kingdom.
Organizational Affiliation: