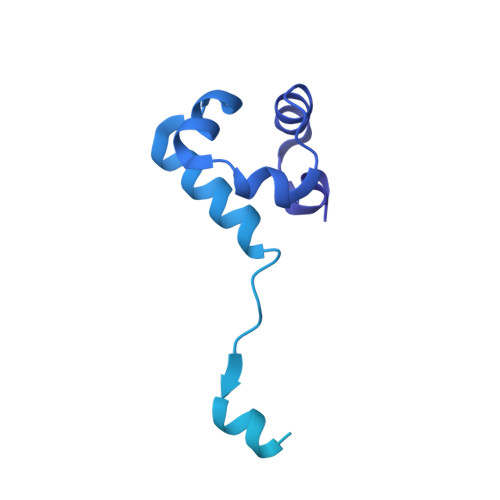
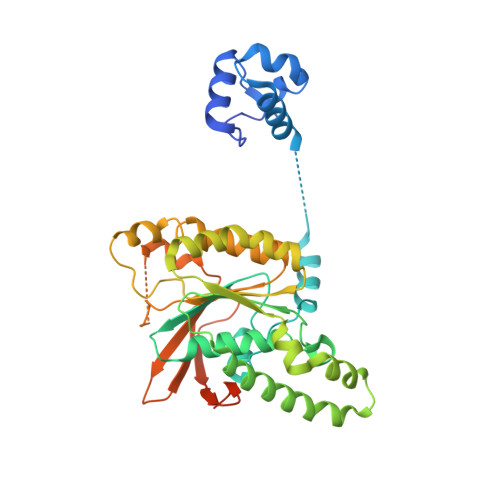
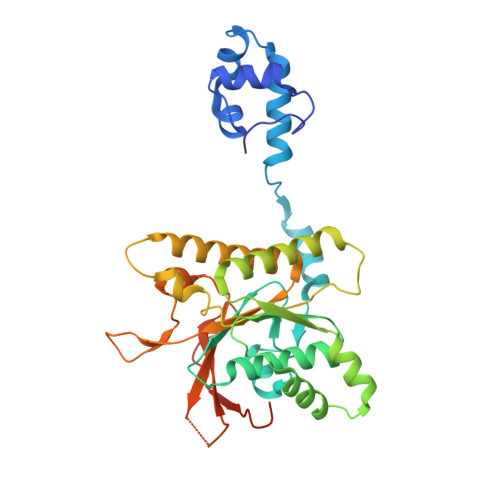
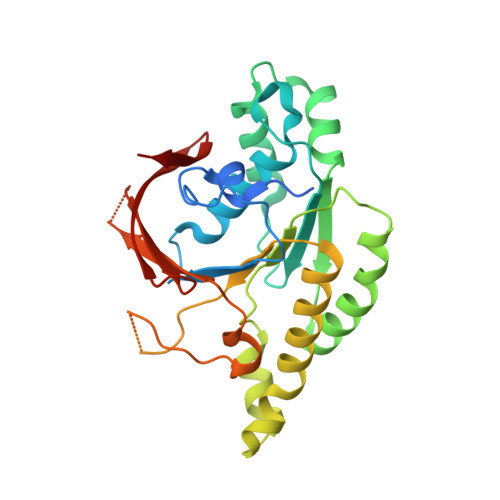
Structure and function of the RAD51B-RAD51C-RAD51D-XRCC2 tumour suppressor.
Greenhough, L.A., Liang, C.C., Belan, O., Kunzelmann, S., Maslen, S., Rodrigo-Brenni, M.C., Anand, R., Skehel, M., Boulton, S.J., West, S.C.(2023) Nature 619: 650-657
- PubMed: 37344587 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-06179-1
- Primary Citation Related Structures:
8OUY, 8OUZ - PubMed Abstract:
Homologous recombination is a fundamental process of life. It is required for the protection and restart of broken replication forks, the repair of chromosome breaks and the exchange of genetic material during meiosis. Individuals with mutations in key recombination genes, such as BRCA2 (also known as FANCD1), or the RAD51 paralogues RAD51B, RAD51C (also known as FANCO), RAD51D, XRCC2 (also known as FANCU) and XRCC3, are predisposed to breast, ovarian and prostate cancers 1-10 and the cancer-prone syndrome Fanconi anaemia 11-13 . The BRCA2 tumour suppressor protein-the product of BRCA2-is well characterized, but the cellular functions of the RAD51 paralogues remain unclear. Genetic knockouts display growth defects, reduced RAD51 focus formation, spontaneous chromosome abnormalities, sensitivity to PARP inhibitors and replication fork defects 14,15 , but the precise molecular roles of RAD51 paralogues in fork stability, DNA repair and cancer avoidance remain unknown. Here we used cryo-electron microscopy, AlphaFold2 modelling and structural proteomics to determine the structure of the RAD51B-RAD51C-RAD51D-XRCC2 complex (BCDX2), revealing that RAD51C-RAD51D-XRCC2 mimics three RAD51 protomers aligned within a nucleoprotein filament, whereas RAD51B is highly dynamic. Biochemical and single-molecule analyses showed that BCDX2 stimulates the nucleation and extension of RAD51 filaments-which are essential for recombinational DNA repair-in reactions that depend on the coupled ATPase activities of RAD51B and RAD51C. Our studies demonstrate that BCDX2 orchestrates RAD51 assembly on single stranded DNA for replication fork protection and double strand break repair, in reactions that are critical for tumour avoidance.
- The Francis Crick Institute, London, UK.
Organizational Affiliation: