Molecular basis of SARS-CoV-2 immune evasion of NTD neutralizing antibodies.
Hong, Q., Xu, S.Q., Huang, Z., Cong, Y.(2024) Dian Zi Xian Wei Xue Bao
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Spike glycoprotein | A [auth E], D [auth C], E [auth D] | 1,261 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 5 Gene Names: S, 2 | 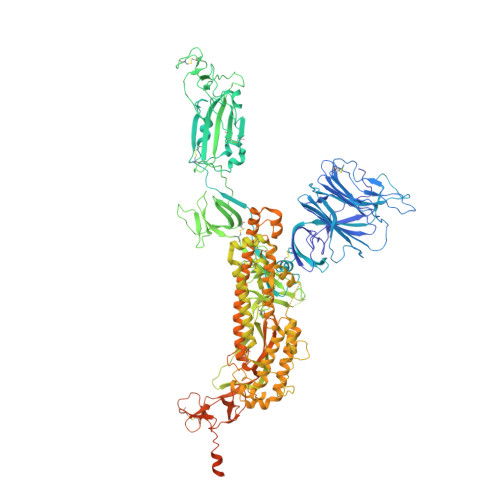 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DTC2 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: P0DTC2-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Heavy chain of S2H5 Fab | B [auth A], F [auth G] | 216 | Mus musculus | Mutation(s): 0 | 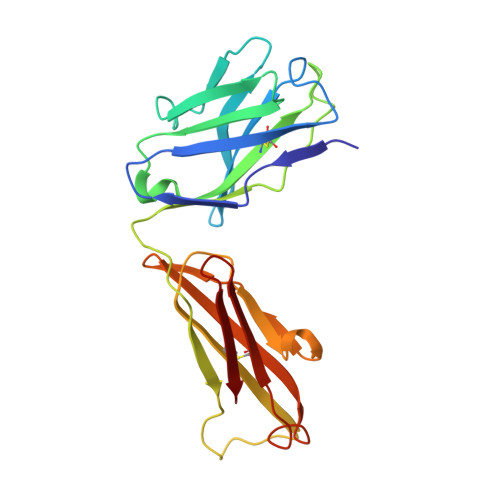 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Light chain of S2H5 Fab | C [auth B], G [auth F] | 219 | Mus musculus | Mutation(s): 0 |  |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | H [auth E] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Chinese Academy of Sciences | China | -- |