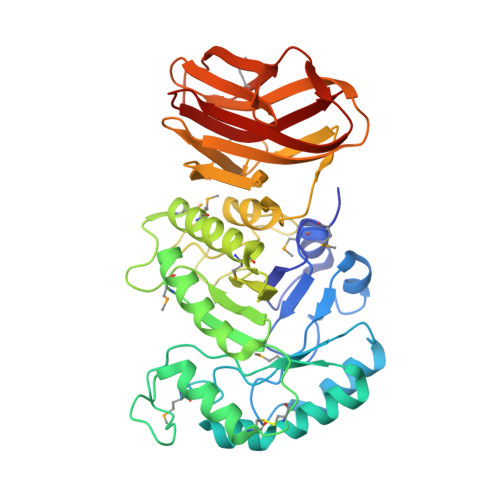
Structural insights into curdlan degradation via a glycoside hydrolase containing a disruptive carbohydrate-binding module.
Lv, T., Feng, J., Jia, X., Wang, C., Li, F., Peng, H., Xiao, Y., Liu, L., He, C.(2024) Biotechnol Biofuels Bioprod 17: 45-45
- PubMed: 38515133 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s13068-024-02494-5
- Primary Citation Related Structures:
8J3X, 8J3Y - PubMed Abstract:
Degradation via enzymatic processes for the production of valuable β-1,3-glucooligosaccharides (GOS) from curdlan has attracted considerable interest. CBM6E functions as a curdlan-specific β-1,3-endoglucanase, composed of a glycoside hydrolase family 128 (GH128) module and a carbohydrate-binding module (CBM) derived from family CBM6. Crystallographic analyses were conducted to comprehend the substrate specificity mechanism of CBM6E. This unveiled structures of both apo CBM6E and its GOS-complexed form. The GH128 and CBM6 modules constitute a cohesive unit, binding nine glucoside moieties within the catalytic groove in a singular helical conformation. By extending the substrate-binding groove, we engineered CBM6E variants with heightened hydrolytic activities, generating diverse GOS profiles from curdlan. Molecular docking, followed by mutation validation, unveiled the cooperative recognition of triple-helical β-1,3-glucan by the GH128 and CBM6 modules, along with the identification of a novel sugar-binding residue situated within the CBM6 module. Interestingly, supplementing the CBM6 module into curdlan gel disrupted the gel's network structure, enhancing the hydrolysis of curdlan by specific β-1,3-glucanases. This study offers new insights into the recognition mechanism of glycoside hydrolases toward triple-helical β-1,3-glucans, presenting an effective method to enhance endoglucanase activity and manipulate its product profile. Furthermore, it discovered a CBM module capable of disrupting the quaternary structures of curdlan, thereby boosting the hydrolytic activity of curdlan gel when co-incubated with β-1,3-glucanases. These findings hold relevance for developing future enzyme and CBM cocktails useful in GOS production from curdlan degradation.
- School of Life Sciences and Anhui Key Laboratory of Modern Biomanufacturing, Anhui University, Hefei, Anhui, China.
Organizational Affiliation: