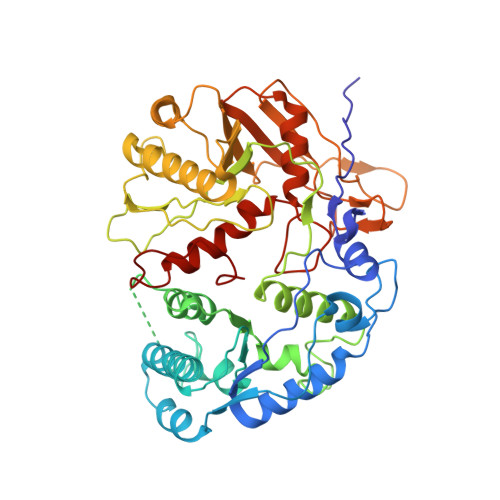
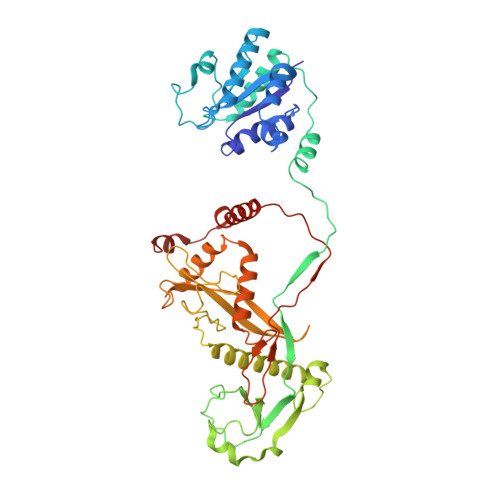
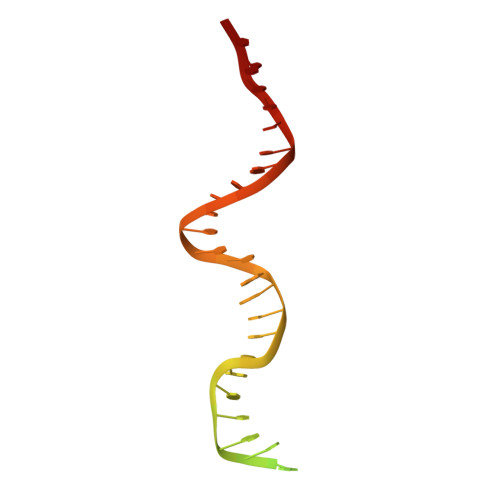
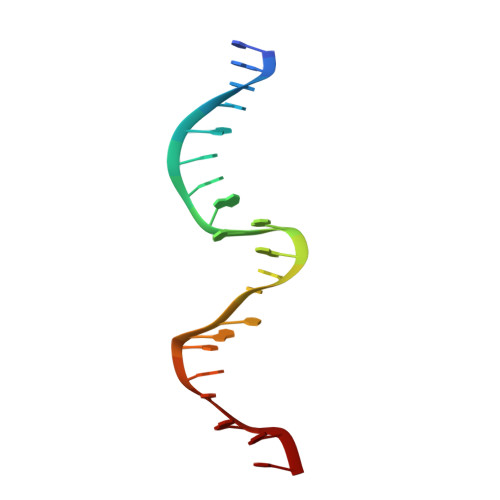
Nucleic-acid-triggered NADase activation of a short prokaryotic Argonaute.
Gao, X., Shang, K., Zhu, K., Wang, L., Mu, Z., Fu, X., Yu, X., Qin, B., Zhu, H., Ding, W., Cui, S.(2024) Nature 625: 822-831
- PubMed: 37783228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-06665-6
- Primary Citation Related Structures:
8ISY, 8ISZ, 8IT0, 8IT1, 8K9G - PubMed Abstract:
Argonaute (Ago) proteins mediate RNA- or DNA-guided inhibition of nucleic acids 1,2 . Although the mechanisms used by eukaryotic Ago proteins and long prokaryotic Ago proteins (pAgos) are known, that used by short pAgos remains elusive. Here we determined the cryo-electron microscopy structures of a short pAgo and the associated TIR-APAZ proteins (SPARTA) from Crenotalea thermophila (Crt): a free-state Crt-SPARTA; a guide RNA-target DNA-loaded Crt-SPARTA; two Crt-SPARTA dimers with distinct TIR organization; and a Crt-SPARTA tetramer. These structures reveal that Crt-SPARTA is composed of a bilobal-fold Ago lobe that connects with a TIR lobe. Whereas the Crt-Ago contains a MID and a PIWI domain, Crt-TIR-APAZ has a TIR domain, an N-like domain, a linker domain and a trigger domain. The bound RNA-DNA duplex adopts a B-form conformation that is recognized by base-specific contacts. Nucleic acid binding causes conformational changes because the trigger domain acts as a 'roadblock' that prevents the guide RNA 5' ends and the target DNA 3' ends from reaching their canonical pockets; this disorders the MID domain and promotes Crt-SPARTA dimerization. Two RNA-DNA-loaded Crt-SPARTA dimers form a tetramer through their TIR domains. Four Crt-TIR domains assemble into two parallel head-to-tail-organized TIR dimers, indicating an NADase-active conformation, which is supported by our mutagenesis study. Our results reveal the structural basis of short-pAgo-mediated defence against invading nucleic acids, and provide insights for optimizing the detection of SPARTA-based programmable DNA sequences.
- NHC Key Laboratory of Systems Biology of Pathogens, Institute of Pathogen Biology, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
Organizational Affiliation: