Structure basis of two nanobodies neutralizing SARS-CoV-2 Omicron variant by targeting ultra-conservative epitopes.
Sun, Z., Wang, L., Li, L., Sun, Y., Zhang, D., Zhou, S., Li, Y., Li, X., Qiao, H., Cui, Q., Lan, Z., Meng, X., Xu, J., Geng, Y., Dai, Y.(2023) J Struct Biol 215: 107996-107996
- PubMed: 37419228 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2023.107996
- Primary Citation Related Structures:
8HR2 - PubMed Abstract:
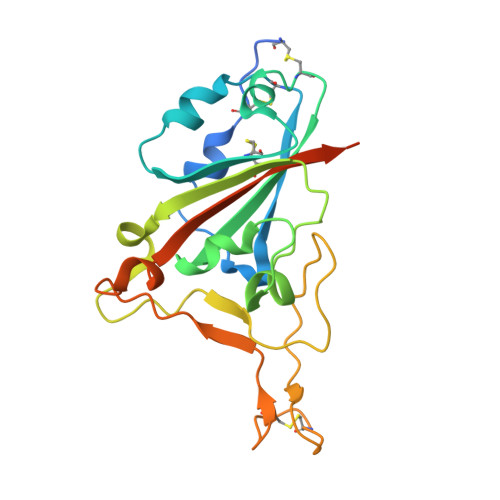
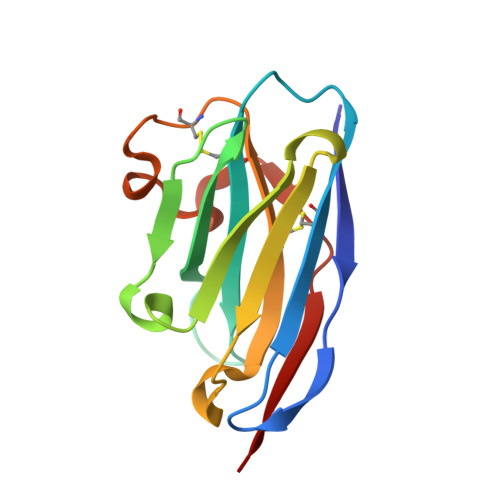
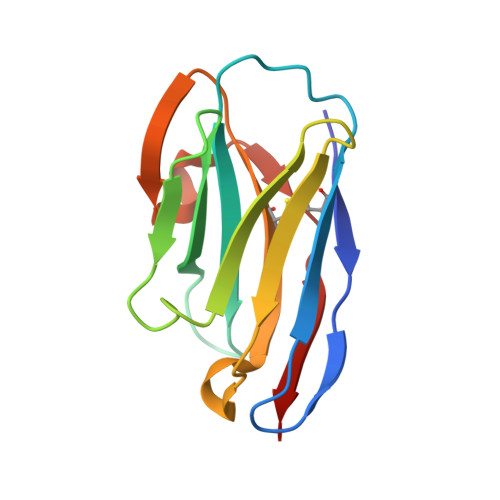
The evolving SARS-CoV-2 Omicron strain has repeatedly caused widespread disease epidemics, and effective antibody drugs continue to be in short supply. Here, we identified a batch of nanobodies with high affinity for receptor binding domain (RBD) of SARS-CoV-2 spike protein, separated them into three classes using high performance liquid chromatography (HPLC), and then resolved the crystal structure of the ternary complexes of two non-competing nanobodies (NB1C6 and NB1B5) with RBD using X-ray crystallography. The structures showed that NB1B5 and NB1C6 bind to the left and right flank of the RBD, respectively, and that the binding epitopes are highly conserved cryptic sites in all SARS-CoV-2 mutant strains, as well as that NB1B5 can effectively block the ACE2. These two nanobodies were covalently linked into multivalent and bi-paratopic formats, and have a high affinity and neutralization potency for omicron, potentially inhibiting viral escape. The binding sites of these two nanobodies are relatively conserved, which help guide the structural design of antibodies targeting future variants of SARS-CoV-2 to combat COVID-19 epidemics and pandemics.
- Department of Biopharmaceutics, College of Food Science and Technology, Shanghai Ocean University, Shanghai 201306, China; State Key Laboratory of Drug Research, The CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China.
Organizational Affiliation: