Structural Basis of the Mechanisms of Action and Immunity of Lactococcin A, a Class IId Bacteriocin.
Li, R., Duan, J., Zhou, Y., Wang, J.(2023) Appl Environ Microbiol 89: e0006623-e0006623
- PubMed: 36840592 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/aem.00066-23
- Primary Citation Related Structures:
8HFS - PubMed Abstract:
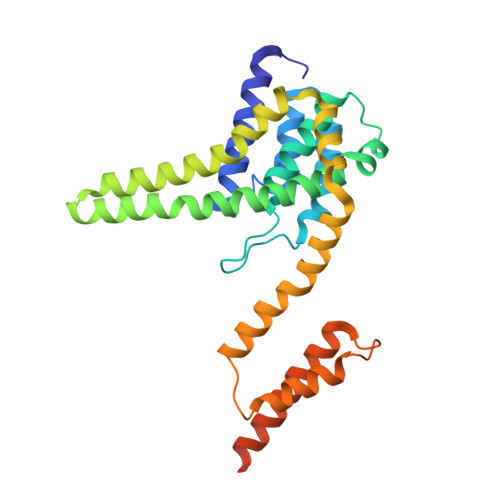
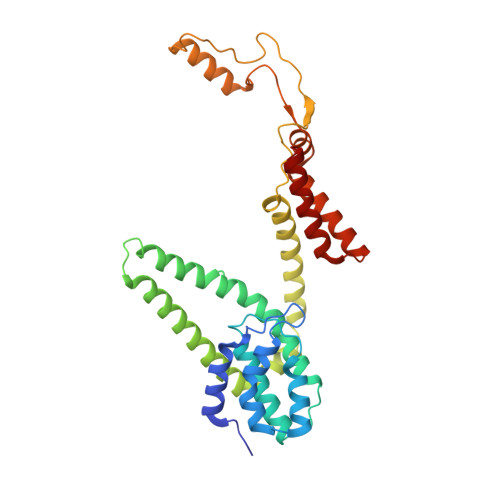
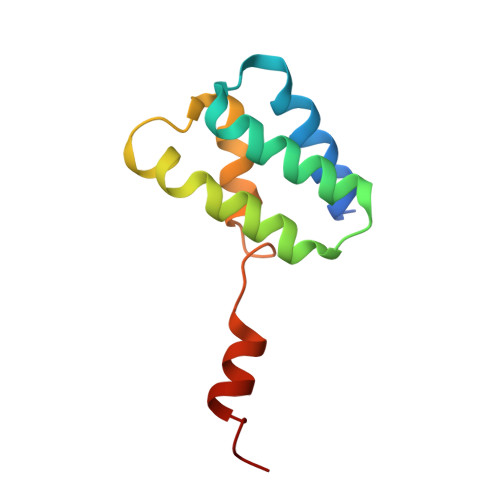
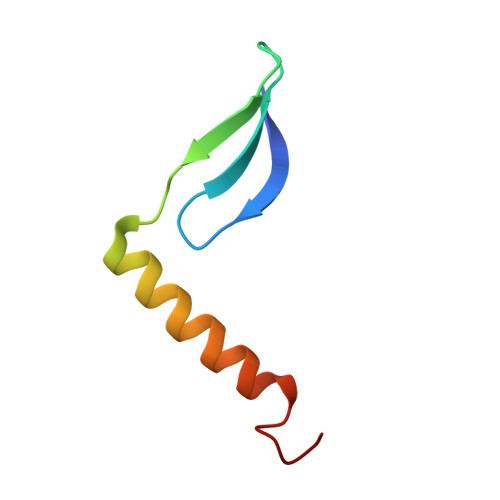
Lactococcin A (LcnA), a class IId bacteriocin, induces membrane leakage and cell death by specifically binding to the membrane receptor-mannose phosphotransferase system (man-PTS), as is the case for pediocin-like (class IIa) bacteriocins. The cognate immunity protein of bacteriocins, which protects the producer cell from its own bacteriocin, recognizes and binds to the bacteriocin-man-PTS complex, consequently blocking membrane leakage. We previously deciphered the mode of action and immunity of class IIa bacteriocins. Here, we determined the structure of the ternary complex of LcnA, LciA ( i.e. , the immunity protein), and its receptor, i.e. , the man-PTS of Lactococcus lactis (ll-man-PTS). An external loop on the membrane-located component IIC of ll-man-PTS was found to prevent specific binding of the N-terminal region of LcnA to the site recognized by pediocin-like bacteriocins. Thus, the N-terminal β-sheet region of LcnA recognized an adjacent site on the extracellular side of ll-man-PTS, with the LcnA C-terminal hydrophobic helix penetrating into the membrane. The cytoplasmic cleft formed within the man-PTS Core and Vmotif domains induced by embedded LcnA from the periplasmic side is adopted by the appropriate angle between helices H3 and H4 of the N terminus of LciA. The flexible C terminus of LciA then blocks membrane leakage. To summarize, our findings reveal the molecular mechanisms of action and immunity of LcnA and LciA, laying a foundation for further design of class IId bacteriocins. IMPORTANCE Class IId (lactococcin-like) bacteriocins and class IIa (pediocin-like) bacteriocins share a few similarities: (i) both induce membrane leakage and cell death by specifically binding the mannose phosphotransferase system (man-PTS) on their target cells, and (ii) cognate immunity proteins recognize and bind to the bacteriocin-man-PTS complex to block membrane leakage. However, class IId bacteriocins lack the "pediocin box" motif, which is typical of class IIa bacteriocins, and basically target only lactococcal cells; in contrast, class IIa bacteriocins target diverse bacterial cells, but not lactococcal cells. We previously solved the structure of class IIa bacteriocin-receptor-immunity ternary complex from Lactobacillus sakei. Here, we determined the structure of the ternary complex of class IId bacteriocin LcnA, its cognate immunity protein LciA, and its receptor, the man-PTS of Lactococcus lactis. By comparing the interactions between man-PTS and class IIa and class IId bacteriocins, this study affords some clues to better understand the specificity of bacteriocins targeting the mannose phosphotransferase system.
- State Key Laboratory of Membrane Biology, Beijing Frontier Research Center for Biological Structure, School of Life Sciences, Tsinghua University, Beijing, People's Republic of China.
Organizational Affiliation: