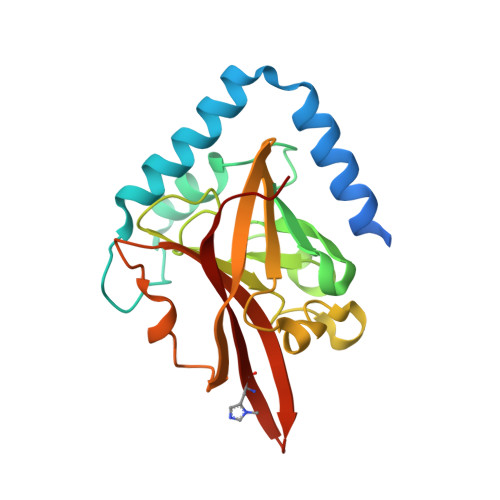
Crystal structure of the pilus-specific sortase from early colonizing oral Streptococcus sanguinis captures an active open-lid conformation.
Yadav, S., Parijat, P., Krishnan, V.(2023) Int J Biol Macromol 243: 125183-125183
- PubMed: 37276901 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2023.125183
- Primary Citation Related Structures:
8GR6, 8GUU - PubMed Abstract:
Dental plaque is a complex microbial biofilm community of many species and a major cause of oral infections and infectious endocarditis. Plaque development begins when primary colonizers attach to oral tissues and undergo coaggregation. Primary colonizers facilitate cellular attachment and inter-bacterial interactions through sortase-dependent pili (or fimbriae) extending out from their cell surface. Consequently, the sortase enzyme is viewed as a potential drug target for controlling biofilm formation and avoiding infection. Streptococcus sanguinis is a primary colonizing bacterium whose pili consist of three different pilin subunits that are assembled together by the pilus-specific (C-type) SsaSrtC sortase. Here, we report on the crystal structure determination of the recombinant wild-type and active-site mutant forms of SsaSrtC. Interestingly, the SsaSrtC structure exhibits an open-lid conformation, although a conserved DPX motif is lacking in the lid. Based on molecular docking and structural analysis, we identified the substrate-binding residues essential for pilin recognition and pilus assembly. We also demonstrated that while recombinant SsaSrtC is enzymatically active toward the five-residue LPNTG sorting motif peptide of the pilins, this activity is significantly reduced by the presence of zinc. We further showed that rutin and α-crocin are potential candidate inhibitors of the SsaSrtC sortase via structure-based virtual screening and inhibition assays. The structural knowledge gained from our study will provide the means to develop new approaches that target pilus-mediated attachment, thereby preventing oral biofilm growth and infection.
- Laboratory of Structural Biology, Regional Centre for Biotechnology, NCR Biotech Science Cluster, Faridabad 121001, India.
Organizational Affiliation: