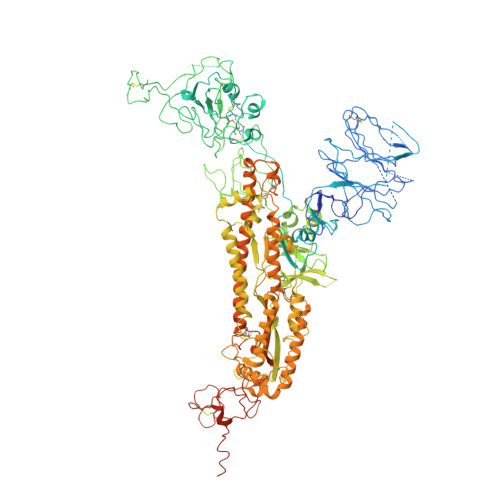
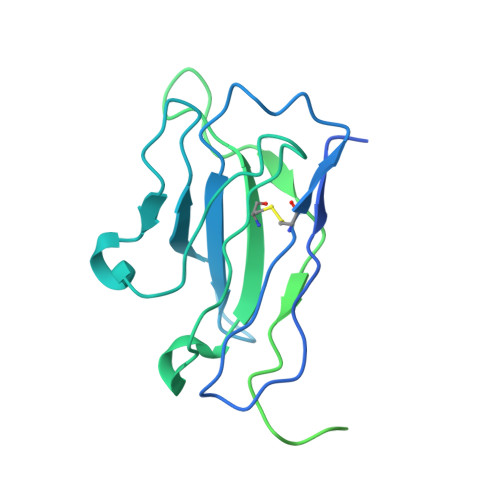
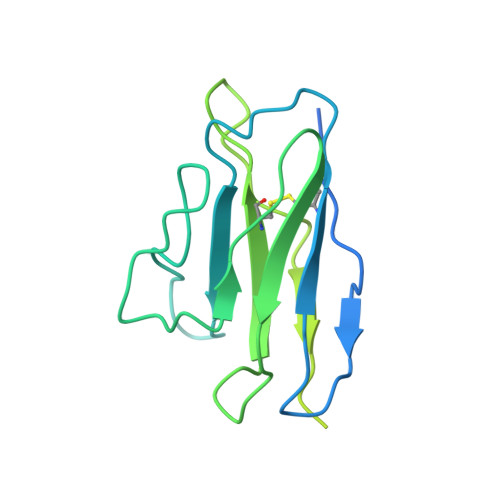
Human antibody BD-218 has broad neutralizing activity against concerning variants of SARS-CoV-2.
Wang, B., Xu, H., Liang, Z.T., Zhao, T.N., Zhang, X., Peng, T.B., Wang, Y.C., Su, X.D.(2023) Int J Biol Macromol 227: 896-902
- PubMed: 36528147 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ijbiomac.2022.12.120
- Primary Citation Related Structures:
8GNH - PubMed Abstract:
As SARS-CoV-2 variants of concern (VOC) reduce the effectiveness of existing anti-COVID therapeutics, it is increasingly critical to identify highly potent neutralizing antibodies (nAbs) that bind to conserved regions across multiple variants, especially beta, delta, and omicron variants. Using single-cell sequencing with biochemical methods and pseudo-typed virus neutralization experiments, here we report the characterization of a potent nAb BD-218, identified from an early screen of patients recovering from the original virus. We have determined the cryo-EM structure of the BD-218/spike protein complex to define its epitope in detail, which revealed that BD-218 interacts with a novel epitope on the receptor-binding domain (RBD) of the spike protein. We concluded that BD-218 is a highly effective and broadly active nAb against SARS-CoV-2 variants with promising potential for therapeutic development.
- State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Biomedical Pioneering Innovation Center (BIOPIC), Peking University, Beijing, China.
Organizational Affiliation: