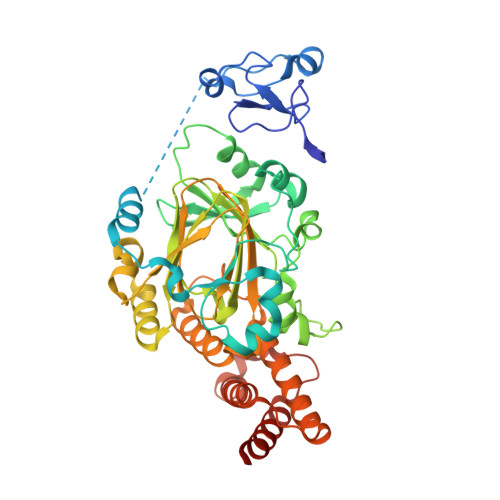
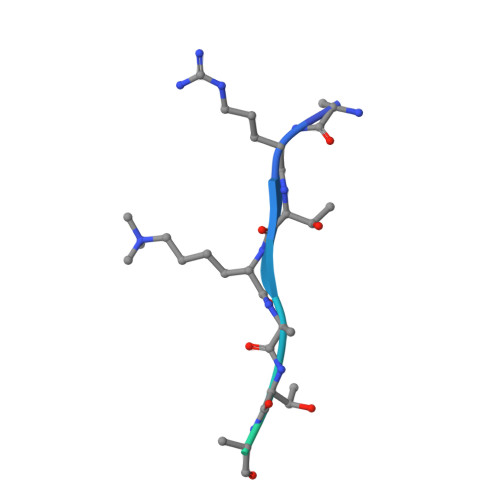
A complete methyl-lysine binding aromatic cage constructed by two domains of PHF2.
Horton, J.R., Zhou, J., Chen, Q., Zhang, X., Bedford, M.T., Cheng, X.(2022) J Biological Chem 299: 102862-102862
- PubMed: 36596360 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.102862
- Primary Citation Related Structures:
8F8Y, 8F8Z - PubMed Abstract:
The N-terminal half of PHF2 harbors both a plant homeodomain (PHD) and a Jumonji domain. The PHD recognizes both histone H3 trimethylated at lysine 4 and methylated nonhistone proteins including vaccinia-related kinase 1 (VRK1). The Jumonji domain erases the repressive dimethylation mark from histone H3 lysine 9 (H3K9me2) at select promoters. The N-terminal amino acid sequences of H3 (AR 2 TK 4 ) and VRK1 (PR 2 VK 4 ) bear an arginine at position 2 and lysine at position 4. Here, we show that the PHF2 N-terminal half binds to H3 and VRK1 peptides containing K4me3, with dissociation constants (K D values) of 160 nM and 42 nM, respectively, which are 4 × and 21 × lower (and higher affinities) than for the isolated PHD domain of PHF2. X-ray crystallography revealed that the K4me3-containing peptide is positioned within the PHD and Jumonji interface, with the positively charged R2 residue engaging acidic residues of the PHD and Jumonji domains and with the K4me3 moiety encircled by aromatic residues from both domains. We suggest that the micromolar binding affinities commonly observed for isolated methyl-lysine reader domains could be improved via additional functional interactions within the same polypeptide or its binding partners.
- Department of Epigenetics and Molecular Carcinogenesis, University of Texas MD Anderson Cancer Center, Houston, Texas, USA.
Organizational Affiliation: