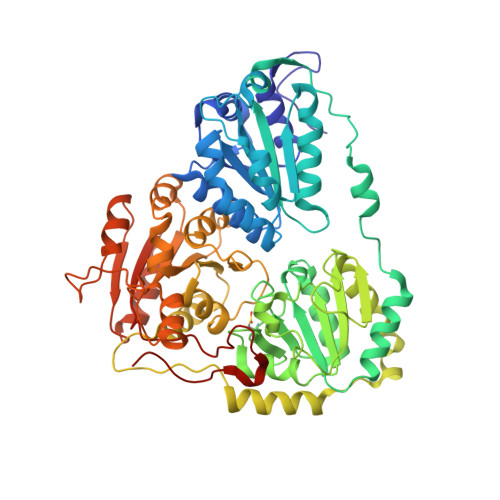
Crystal Structure of the Commercial Herbicide, Amidosulfuron, in Complex with Arabidopsis thaliana Acetohydroxyacid Synthase.
Cheng, Y., Lonhienne, T., Garcia, M.D., Williams, C.M., Schenk, G., Guddat, L.W.(2023) J Agric Food Chem 71: 5117-5126
- PubMed: 36943718 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jafc.2c08528
- Primary Citation Related Structures:
8ET4, 8ET5 - PubMed Abstract:
Amidosulfuron (AS) is from the commercial sulfonylurea herbicide family. It is highly effective against dicot broad-leaf weeds. This herbicide targets acetohydroxyacid synthase (AHAS), the first enzyme in the branched chain amino acid biosynthesis pathway. Here, we have determined the crystal structure of AS in complex with wildtype Arabidopsis thaliana AHAS ( At AHAS) and with the resistance mutant, S653T. In both structures, the cofactor, ThDP, is modified to a peracetate adduct, consistent with time-dependent accumulative inhibition. Compared to other AHAS-inhibiting herbicides of the sulfonylurea family, AS lacks a second aromatic ring. The replacement is an aryl sulfonyl group with a reduced number of interactions with the enzyme and relatively low affinity ( K i = 4.2 μM vs low nM when two heteroaromatic rings are present). This study shows that effective herbicides can have a relatively high K i for plant AHAS but can still be a potent herbicide provided accumulative inhibition also occurs.
- School of Chemistry and Molecular Biosciences, The University of Queensland, Brisbane 4072, Australia.
Organizational Affiliation: