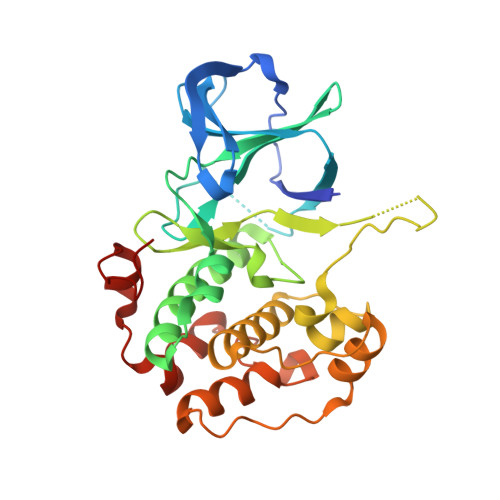
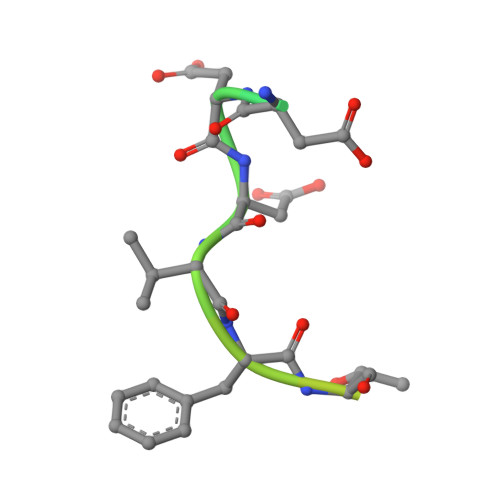
The ribosomal S6 kinase 2 (RSK2)-SPRED2 complex regulates the phosphorylation of RSK substrates and MAPK signaling.
Lopez, J., Bonsor, D.A., Sale, M.J., Urisman, A., Mehalko, J.L., Cabanski-Dunning, M., Castel, P., Simanshu, D.K., McCormick, F.(2023) J Biological Chem 299: 104789-104789
- PubMed: 37149146 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.104789
- Primary Citation Related Structures:
8EQ5 - PubMed Abstract:
Sprouty-related EVH-1 domain-containing (SPRED) proteins are a family of proteins that negatively regulate the RAS-Mitogen-Activated Protein Kinase (MAPK) pathway, which is involved in the regulation of the mitogenic response and cell proliferation. However, the mechanism by which these proteins affect RAS-MAPK signaling has not been elucidated. Patients with mutations in SPRED give rise to unique disease phenotypes; thus, we hypothesized that distinct interactions across SPRED proteins may account for alternative nodes of regulation. To characterize the SPRED interactome and evaluate how members of the SPRED family function through unique binding partners, we performed affinity purification mass spectrometry. We identified 90-kDa ribosomal S6 kinase 2 (RSK2) as a specific interactor of SPRED2 but not SPRED1 or SPRED3. We identified that the N-terminal kinase domain of RSK2 mediates the interaction between amino acids 123 to 201 of SPRED2. Using X-ray crystallography, we determined the structure of the SPRED2-RSK2 complex and identified the SPRED2 motif, F145A, as critical for interaction. We found that the formation of this interaction is regulated by MAPK signaling events. We also find that this interaction between SPRED2 and RSK2 has functional consequences, whereby the knockdown of SPRED2 resulted in increased phosphorylation of RSK substrates, YB1 and CREB. Furthermore, SPRED2 knockdown hindered phospho-RSK membrane and nuclear subcellular localization. We report that disruption of the SPRED2-RSK complex has effects on RAS-MAPK signaling dynamics. Our analysis reveals that members of the SPRED family have unique protein binding partners and describes the molecular and functional determinants of SPRED2-RSK2 complex dynamics.
- Helen Diller Family Comprehensive Cancer Center, University of California, San Francisco, San Francisco, California, USA.
Organizational Affiliation: