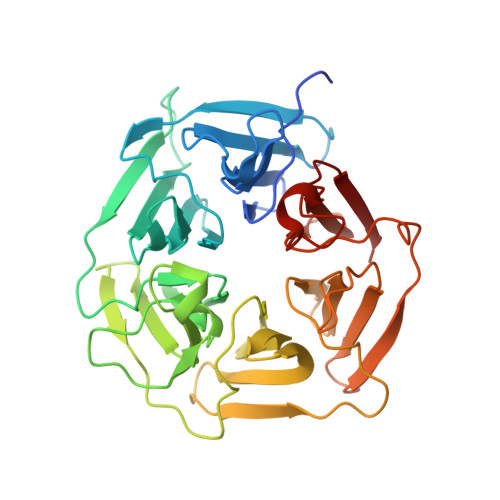
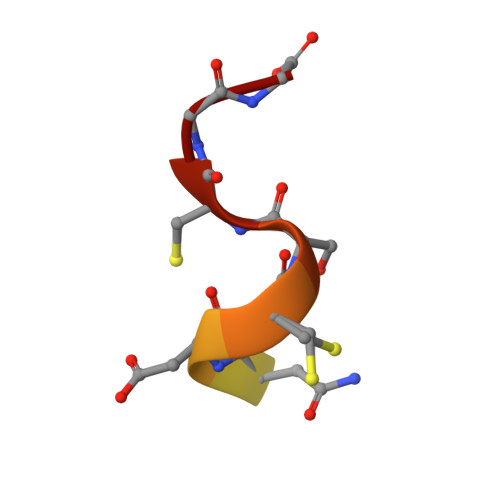
E3 ligase autoinhibition by C-degron mimicry maintains C-degron substrate fidelity.
Scott, D.C., King, M.T., Baek, K., Gee, C.T., Kalathur, R., Li, J., Purser, N., Nourse, A., Chai, S.C., Vaithiyalingam, S., Chen, T., Lee, R.E., Elledge, S.J., Kleiger, G., Schulman, B.A.(2023) Mol Cell 83: 770-786.e9
- PubMed: 36805027 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2023.01.019
- Primary Citation Related Structures:
8EBL, 8EBM, 8EBN - PubMed Abstract:
E3 ligase recruitment of proteins containing terminal destabilizing motifs (degrons) is emerging as a major form of regulation. How those E3s discriminate bona fide substrates from other proteins with terminal degron-like sequences remains unclear. Here, we report that human KLHDC2, a CRL2 substrate receptor targeting C-terminal Gly-Gly degrons, is regulated through interconversion between two assemblies. In the self-inactivated homotetramer, KLHDC2's C-terminal Gly-Ser motif mimics a degron and engages the substrate-binding domain of another protomer. True substrates capture the monomeric CRL2 KLHDC2 , driving E3 activation by neddylation and subsequent substrate ubiquitylation. Non-substrates such as NEDD8 bind KLHDC2 with high affinity, but its slow on rate prevents productive association with CRL2 KLHDC2 . Without substrate, neddylated CRL2 KLHDC2 assemblies are deactivated via distinct mechanisms: the monomer by deneddylation and the tetramer by auto-ubiquitylation. Thus, substrate specificity is amplified by KLHDC2 self-assembly acting like a molecular timer, where only bona fide substrates may bind before E3 ligase inactivation.
- Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, TN 38105, USA. Electronic address: danny.scott@stjude.org.
Organizational Affiliation: