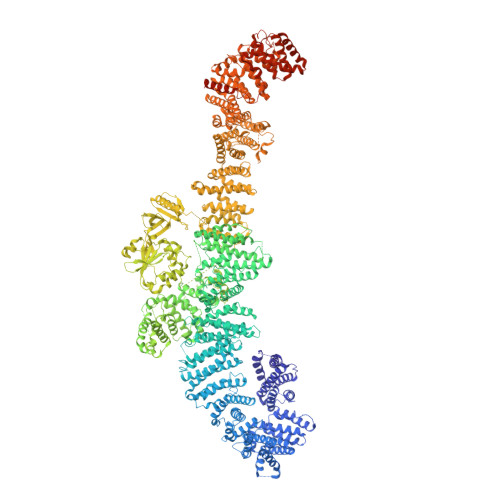
Destabilizing NF1 variants act in a dominant negative manner through neurofibromin dimerization.
Young, L.C., Goldstein de Salazar, R., Han, S.W., Huang, Z.Y.S., Merk, A., Drew, M., Darling, J., Wall, V., Grisshammer, R., Cheng, A., Allison, M.R., Sale, M.J., Nissley, D.V., Esposito, D., Ognjenovic, J., McCormick, F.(2023) Proc Natl Acad Sci U S A 120: e2208960120-e2208960120
- PubMed: 36689660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2208960120
- Primary Citation Related Structures:
8E20, 8EDM - PubMed Abstract:
The majority of pathogenic mutations in the neurofibromatosis type I ( NF1 ) gene reduce total neurofibromin protein expression through premature truncation or microdeletion, but it is less well understood how loss-of-function missense variants drive NF1 disease. We have found that patient variants in codons 844 to 848, which correlate with a severe phenotype, cause protein instability and exert an additional dominant-negative action whereby wild-type neurofibromin also becomes destabilized through protein dimerization. We have used our neurofibromin cryogenic electron microscopy structure to predict and validate other patient variants that act through a similar mechanism. This provides a foundation for understanding genotype-phenotype correlations and has important implications for patient counseling, disease management, and therapeutics.
- Helen Diller Family Comprehensive Cancer Center, University of California, San Francisco, CA 94153.
Organizational Affiliation: