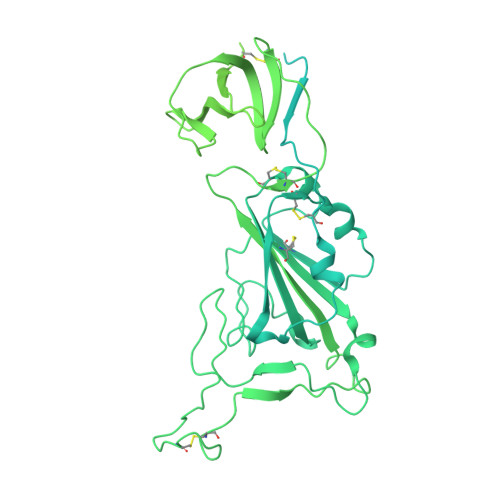
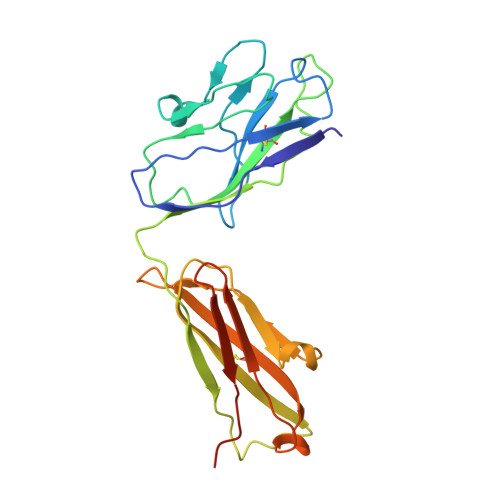
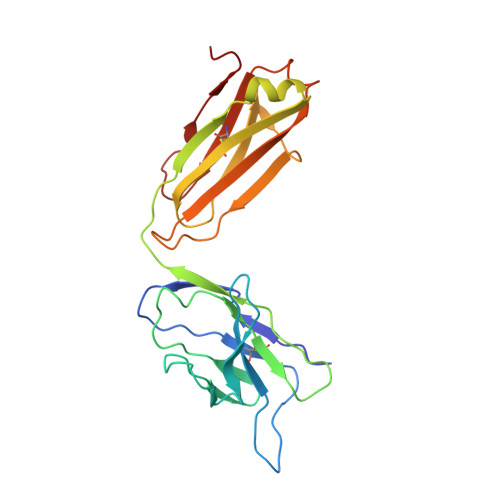
Human neutralizing antibodies to cold linear epitopes and subdomain 1 of the SARS-CoV-2 spike glycoprotein.
Bianchini, F., Crivelli, V., Abernathy, M.E., Guerra, C., Palus, M., Muri, J., Marcotte, H., Piralla, A., Pedotti, M., De Gasparo, R., Simonelli, L., Matkovic, M., Toscano, C., Biggiogero, M., Calvaruso, V., Svoboda, P., Cervantes Rincon, T., Fava, T., Podesvova, L., Shanbhag, A.A., Celoria, A., Sgrignani, J., Stefanik, M., Honig, V., Pranclova, V., Michalcikova, T., Prochazka, J., Guerrini, G., Mehn, D., Ciabattini, A., Abolhassani, H., Jarrossay, D., Uguccioni, M., Medaglini, D., Pan-Hammarstrom, Q., Calzolai, L., Fernandez, D., Baldanti, F., Franzetti-Pellanda, A., Garzoni, C., Sedlacek, R., Ruzek, D., Varani, L., Cavalli, A., Barnes, C.O., Robbiani, D.F.(2023) Sci Immunol 8: eade0958-eade0958
- PubMed: 36701425 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciimmunol.ade0958
- Primary Citation Related Structures:
8D47, 8D48 - PubMed Abstract:
Emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants diminishes the efficacy of vaccines and antiviral monoclonal antibodies. Continued development of immunotherapies and vaccine immunogens resilient to viral evolution is therefore necessary. Using coldspot-guided antibody discovery, a screening approach that focuses on portions of the virus spike glycoprotein that are both functionally relevant and averse to change, we identified human neutralizing antibodies to highly conserved viral epitopes. Antibody fp.006 binds the fusion peptide and cross-reacts against coronaviruses of the four genera, including the nine human coronaviruses, through recognition of a conserved motif that includes the S2' site of proteolytic cleavage. Antibody hr2.016 targets the stem helix and neutralizes SARS-CoV-2 variants. Antibody sd1.040 binds to subdomain 1, synergizes with antibody rbd.042 for neutralization, and, similar to fp.006 and hr2.016, protects mice expressing human angiotensin-converting enzyme 2 against infection when present as a bispecific antibody. Thus, coldspot-guided antibody discovery reveals donor-derived neutralizing antibodies that are cross-reactive with Orthocoronavirinae, including SARS-CoV-2 variants.
- Institute for Research in Biomedicine, Università della Svizzera italiana, Bellinzona, Switzerland.
Organizational Affiliation: