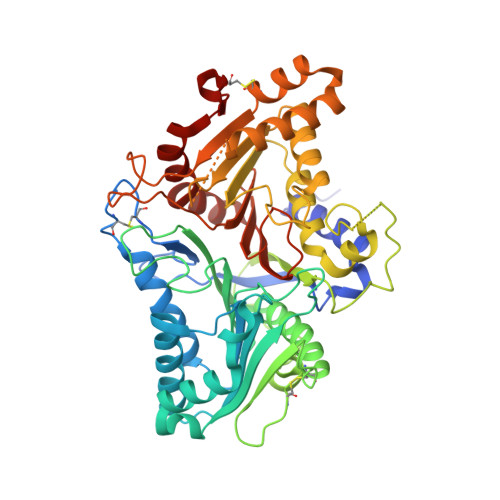
Activity-based directed evolution of a membrane editor in mammalian cells.
Tei, R., Bagde, S.R., Fromme, J.C., Baskin, J.M.(2023) Nat Chem 15: 1030-1039
- PubMed: 37217787 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41557-023-01214-0
- Primary Citation Related Structures:
8CTP, 8CTQ - PubMed Abstract:
Cellular membranes contain numerous lipid species, and efforts to understand the biological functions of individual lipids have been stymied by a lack of approaches for controlled modulation of membrane composition in situ. Here we present a strategy for editing phospholipids, the most abundant lipids in biological membranes. Our membrane editor is based on a bacterial phospholipase D (PLD), which exchanges phospholipid head groups through hydrolysis or transphosphatidylation of phosphatidylcholine with water or exogenous alcohols. Exploiting activity-dependent directed enzyme evolution in mammalian cells, we have developed and structurally characterized a family of 'superPLDs' with up to a 100-fold enhancement in intracellular activity. We demonstrate the utility of superPLDs for both optogenetics-enabled editing of phospholipids within specific organelle membranes in live cells and biocatalytic synthesis of natural and unnatural designer phospholipids in vitro. Beyond the superPLDs, activity-based directed enzyme evolution in mammalian cells is a generalizable approach to engineer additional chemoenzymatic biomolecule editors.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, NY, USA.
Organizational Affiliation: