Structural insights into the broad protection against H1 influenza viruses by a computationally optimized hemagglutinin vaccine.
Dzimianski, J.V., Han, J., Sautto, G.A., O'Rourke, S.M., Cruz, J.M., Pierce, S.R., Ecker, J.W., Carlock, M.A., Nagashima, K.A., Mousa, J.J., Ross, T.M., Ward, A.B., DuBois, R.M.(2023) Commun Biol 6: 454-454
- PubMed: 37185989 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-023-04793-3
- Primary Citation Related Structures:
7UYI, 8CT6 - PubMed Abstract:
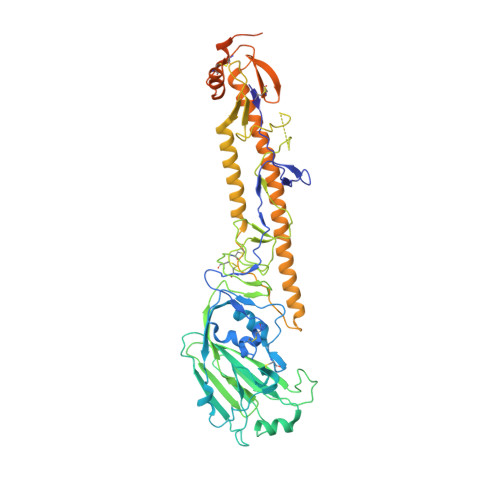
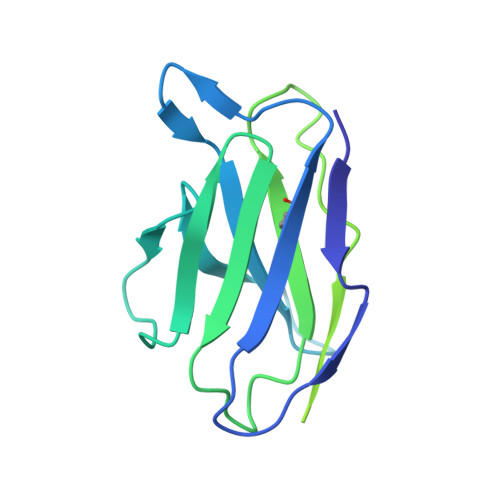
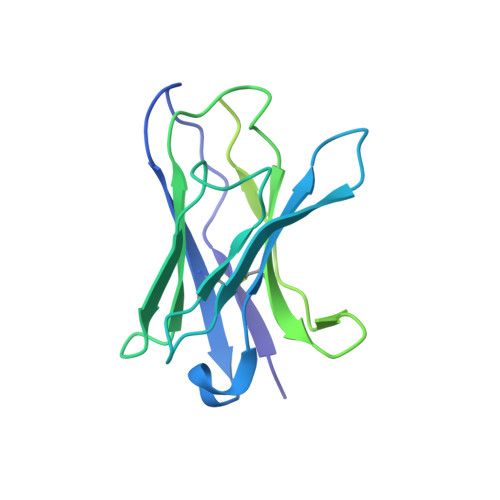
Influenza virus poses an ongoing human health threat with pandemic potential. Due to mutations in circulating strains, formulating effective vaccines remains a challenge. The use of computationally optimized broadly reactive antigen (COBRA) hemagglutinin (HA) proteins is a promising vaccine strategy to protect against a wide range of current and future influenza viruses. Though effective in preclinical studies, the mechanistic basis driving the broad reactivity of COBRA proteins remains to be elucidated. Here, we report the crystal structure of the COBRA HA termed P1 and identify antigenic and glycosylation properties that contribute to its immunogenicity. We further report the cryo-EM structure of the P1-elicited broadly neutralizing antibody 1F8 bound to COBRA P1, revealing 1F8 to recognize an atypical receptor binding site epitope via an unexpected mode of binding.
- Department of Biomolecular Engineering, University of California Santa Cruz, Santa Cruz, CA, USA.
Organizational Affiliation: