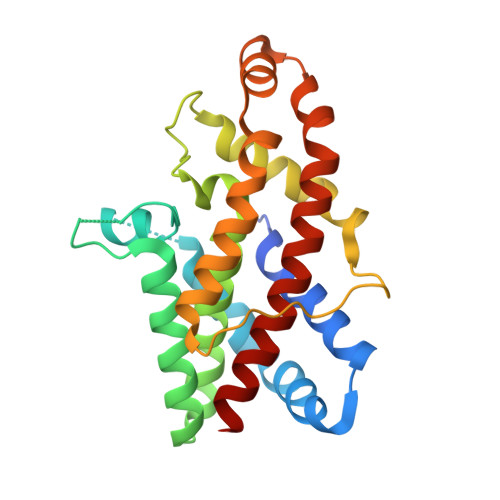
Structural basis for the unique molecular properties of broad-range phospholipase C from Listeria monocytogenes.
Petrisic, N., Adamek, M., Kezar, A., Hocevar, S.B., Zagar, E., Anderluh, G., Podobnik, M.(2023) Nat Commun 14: 6474-6474
- PubMed: 37838694 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-42134-4
- Primary Citation Related Structures:
8CQM - PubMed Abstract:
Listeriosis is one of the most serious foodborne diseases caused by the intracellular bacterium Listeria monocytogenes. Its two major virulence factors, broad-range phospholipase C (LmPC-PLC) and the pore-forming toxin listeriolysin O (LLO), enable the bacterium to spread in the host by destroying cell membranes. Here, we determine the crystal structure of LmPC-PLC and complement it with the functional analysis of this enzyme. This reveals that LmPC-PLC has evolved several structural features to regulate its activity, including the invariant position of the N-terminal tryptophan (W1), the structurally plastic active site, Zn 2+ -dependent activity, and the tendency to form oligomers with impaired enzymatic activity. We demonstrate that the enzymatic activity of LmPC-PLC can be specifically inhibited by its propeptide added in trans. Furthermore, we show that the phospholipase activity of LmPC-PLC facilitates the pore-forming activity of LLO and affects the morphology of LLO oligomerization on lipid membranes, revealing the multifaceted synergy of the two virulence factors.
- Department of Molecular Biology and Nanobiotechnology, National Institute of Chemistry, Ljubljana, Slovenia.
Organizational Affiliation: