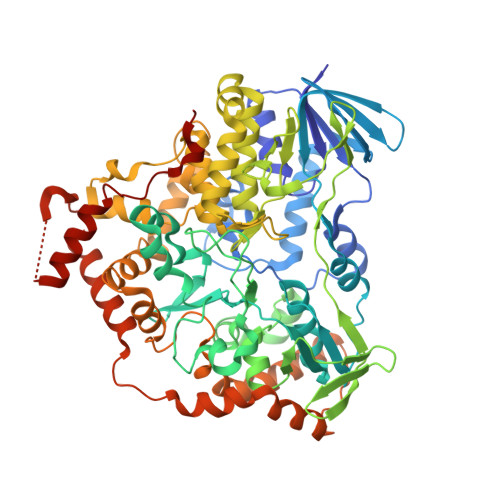
Structural basis of regioselective tryptophan dibromination by the single-component flavin-dependent halogenase AetF.
Gafe, S., Niemann, H.H.(2023) Acta Crystallogr D Struct Biol 79: 596-609
- PubMed: 37314407 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798323004254
- Primary Citation Related Structures:
8CJD, 8CJE, 8CJF, 8CJG - PubMed Abstract:
The flavin-dependent halogenase (FDH) AetF successively brominates tryptophan at C5 and C7 to generate 5,7-dibromotryptophan. In contrast to the well studied two-component tryptophan halogenases, AetF is a single-component flavoprotein monooxygenase. Here, crystal structures of AetF alone and in complex with various substrates are presented, representing the first experimental structures of a single-component FDH. Rotational pseudosymmetry and pseudomerohedral twinning complicated the phasing of one structure. AetF is structurally related to flavin-dependent monooxygenases. It contains two dinucleotide-binding domains for binding the ADP moiety with unusual sequences that deviate from the consensus sequences GXGXXG and GXGXXA. A large domain tightly binds the cofactor flavin adenine dinucleotide (FAD), while the small domain responsible for binding the nicotinamide adenine dinucleotide (NADP) is unoccupied. About half of the protein forms additional structural elements containing the tryptophan binding site. FAD and tryptophan are about 16 Å apart. A tunnel between them presumably allows diffusion of the active halogenating agent hypohalous acid from FAD to the substrate. Tryptophan and 5-bromotryptophan bind to the same site but with a different binding pose. A flip of the indole moiety identically positions C5 of tryptophan and C7 of 5-bromotryptophan next to the tunnel and to catalytic residues, providing a simple explanation for the regioselectivity of the two successive halogenations. AetF can also bind 7-bromotryptophan in the same orientation as tryptophan. This opens the way for the biocatalytic production of differentially dihalogenated tryptophan derivatives. The structural conservation of a catalytic lysine suggests a way to identify novel single-component FDHs.
- Department of Chemistry, Bielefeld University, Universitaetsstrasse 25, 33615 Bielefeld, Germany.
Organizational Affiliation: