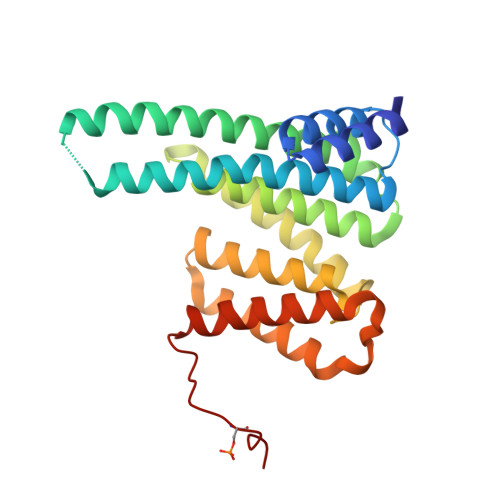
Structural basis for the recognition by 14-3-3 proteins of a conditional binding site within the oligomerization domain of human nucleophosmin.
Kapitonova, A.A., Tugaeva, K.V., Varfolomeeva, L.A., Boyko, K.M., Cooley, R.B., Sluchanko, N.N.(2022) Biochem Biophys Res Commun 627: 176-183
- PubMed: 36041327 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2022.08.047
- Primary Citation Related Structures:
8AH2 - PubMed Abstract:
Nucleophosmin 1 (NPM1) is a multifunctional protein regulating ribosome biogenesis, centrosome duplication and chromatin remodeling. Being a major nucleolar protein, NPM1 can migrate to the nucleus and the cytoplasm, which is controlled by changes of NPM1 oligomerization and interaction with other cell factors. NPM1 forms a stable pentamer with its N-terminal structured domain, where two nuclear export signals and several phosphorylation sites reside. This domain undergoes dissociation and disordering upon Ser48 phosphorylation in the subunit interface. Recent studies indicated that Ser48 is important for NPM1 interaction with other proteins including 14-3-3, the well-known phosphoserine/phosphothreonine binders, but the structural basis for 14-3-3/NPM1 interaction remained unaddressed. By fusing human 14-3-3ζ with an NPM1 segment surrounding Ser48, which was phosphorylated inside Escherichia coli cells by co-expressed protein kinase A, here we obtained the desired protein/phosphopeptide complex and determined its crystal structure. While biochemical data indicated that the interaction is driven by Ser48 phosphorylation, the crystallographic 14-3-3/phosphopeptide interface reveals an NPM1 conformation distinctly different from that in the NPM1 pentamer. Given the canonical phosphopeptide-binding mode observed in our crystal structure, Ser48 emerges as a conditional binding site whose recognition by 14-3-3 proteins is enabled by NPM1 phosphorylation, disassembly and disordering under physiological circumstances.
- A.N. Bach Institute of Biochemistry, Federal Research Center of Biotechnology of the Russian Academy of Sciences, 119071, Moscow, Russia.
Organizational Affiliation: