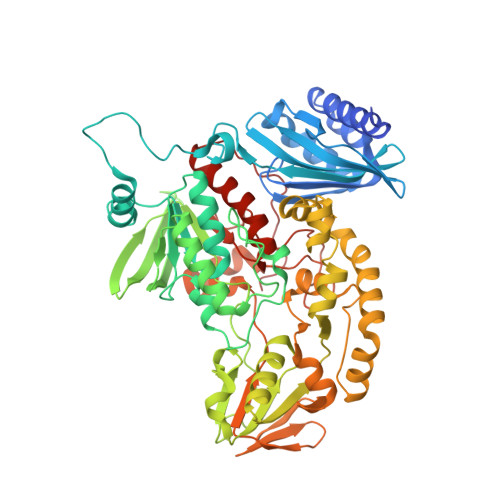
A Cold-Active Flavin-Dependent Monooxygenase from Janthinobacterium svalbardensis Unlocks Applications of Baeyer-Villiger Monooxygenases at Low Temperature.
Chanique, A.M., Polidori, N., Sovic, L., Kracher, D., Assil-Companioni, L., Galuska, P., Parra, L.P., Gruber, K., Kourist, R.(2023) ACS Catal 13: 3549-3562
- PubMed: 36970468 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acscatal.2c05160
- Primary Citation Related Structures:
8ACS - PubMed Abstract:
Cold-active enzymes maintain a large part of their optimal activity at low temperatures. Therefore, they can be used to avoid side reactions and preserve heat-sensitive compounds. Baeyer-Villiger monooxygenases (BVMO) utilize molecular oxygen as a co-substrate to catalyze reactions widely employed for steroid, agrochemical, antibiotic, and pheromone production. Oxygen has been described as the rate-limiting factor for some BVMO applications, thereby hindering their efficient utilization. Considering that oxygen solubility in water increases by 40% when the temperature is decreased from 30 to 10 °C, we set out to identify and characterize a cold-active BVMO. Using genome mining in the Antarctic organism Janthinobacterium svalbardensis, a cold-active type II flavin-dependent monooxygenase (FMO) was discovered. The enzyme shows promiscuity toward NADH and NADPH and high activity between 5 and 25 °C. The enzyme catalyzes the monooxygenation and sulfoxidation of a wide range of ketones and thioesters. The high enantioselectivity in the oxidation of norcamphor (eeS = 56%, eeP > 99%, E > 200) demonstrates that the generally higher flexibility observed in the active sites of cold-active enzymes, which compensates for the lower motion at cold temperatures, does not necessarily reduce the selectivity of these enzymes. To gain a better understanding of the unique mechanistic features of type II FMOs, we determined the structure of the dimeric enzyme at 2.5 Å resolution. While the unusual N-terminal domain has been related to the catalytic properties of type II FMOs, the structure shows a SnoaL-like N-terminal domain that is not interacting directly with the active site. The active site of the enzyme is accessible only through a tunnel, with Tyr-458, Asp-217, and His-216 as catalytic residues, a combination not observed before in FMOs and BVMOs.
- NAWI Graz, BioTechMed-Graz, Institute of Molecular Biotechnology, Graz University of Technology, Petersgasse 14, Graz 8010, Austria.
Organizational Affiliation: