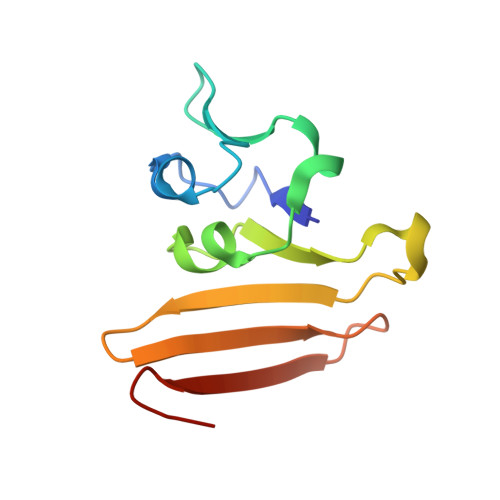
Assimilatory sulfate reduction in the marine methanogen Methanothermococcus thermolithotrophicus.
Jespersen, M., Wagner, T.(2023) Nat Microbiol 8: 1227-1239
- PubMed: 37277534 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-023-01398-8
- Primary Citation Related Structures:
8A8D, 8A8G, 8A8H, 8A8K, 8A8O - PubMed Abstract:
Methanothermococcus thermolithotrophicus is the only known methanogen that grows on sulfate as its sole sulfur source, uniquely uniting methanogenesis and sulfate reduction. Here we use physiological, biochemical and structural analyses to provide a snapshot of the complete sulfate reduction pathway of this methanogenic archaeon. We find that later steps in this pathway are catalysed by atypical enzymes. PAPS (3'-phosphoadenosine 5'-phosphosulfate) released by APS kinase is converted into sulfite and 3'-phosphoadenosine 5'-phosphate (PAP) by a PAPS reductase that is similar to the APS reductases of dissimilatory sulfate reduction. A non-canonical PAP phosphatase then hydrolyses PAP. Finally, the F 420 -dependent sulfite reductase converts sulfite to sulfide for cellular assimilation. While metagenomic and metatranscriptomic studies suggest that the sulfate reduction pathway is present in several methanogens, the sulfate assimilation pathway in M. thermolithotrophicus is distinct. We propose that this pathway was 'mix-and-matched' through the acquisition of assimilatory and dissimilatory enzymes from other microorganisms and then repurposed to fill a unique metabolic role.
- Microbial Metabolism Group, Max Planck Institute for Marine Microbiology, Bremen, Germany.
Organizational Affiliation: