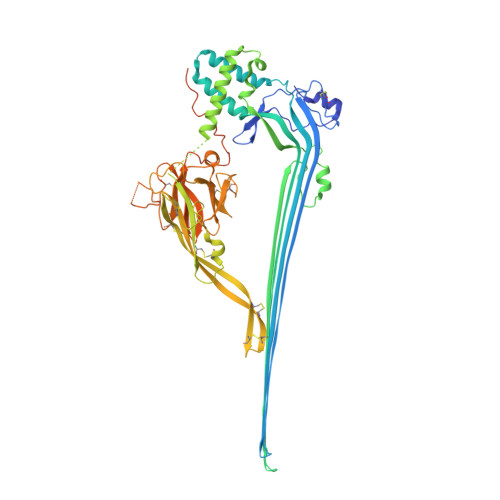
Cryo-EM structures of perforin-2 in isolation and assembled on a membrane suggest a mechanism for pore formation.
Yu, X., Ni, T., Munson, G., Zhang, P., Gilbert, R.J.C.(2022) EMBO J 41: e111857-e111857
- PubMed: 36245269 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2022111857
- Primary Citation Related Structures:
8A1D, 8A1S - PubMed Abstract:
Perforin-2 (PFN2, MPEG1) is a key pore-forming protein in mammalian innate immunity restricting intracellular bacteria proliferation. It forms a membrane-bound pre-pore complex that converts to a pore-forming structure upon acidification; but its mechanism of conformational transition has been debated. Here we used cryo-electron microscopy, tomography and subtomogram averaging to determine structures of PFN2 in pre-pore and pore conformations in isolation and bound to liposomes. In isolation and upon acidification, the pre-assembled complete pre-pore rings convert to pores in both flat ring and twisted conformations. On membranes, in situ assembled PFN2 pre-pores display various degrees of completeness; whereas PFN2 pores are mainly incomplete arc structures that follow the same subunit packing arrangements as found in isolation. Both assemblies on membranes use their P2 β-hairpin for binding to the lipid membrane surface. Overall, these structural snapshots suggest a molecular mechanism for PFN2 pre-pore to pore transition on a targeted membrane, potentially using the twisted pore as an intermediate or alternative state to the flat conformation, with the capacity to cause bilayer distortion during membrane insertion.
- Division of Structural Biology, Wellcome Centre for Human Genetics, University of Oxford, Oxford, UK.
Organizational Affiliation: