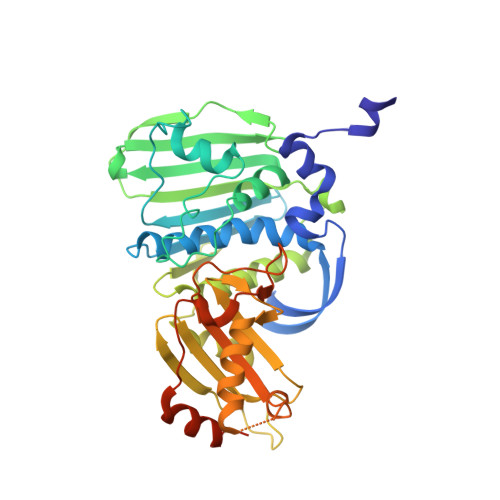
A comprehensive structural analysis of the ATPase domain of human DNA topoisomerase II beta bound to AMPPNP, ADP, and the bisdioxopiperazine, ICRF193.
Ling, E.M., Basle, A., Cowell, I.G., van den Berg, B., Blower, T.R., Austin, C.A.(2022) Structure 30: 1129-1145.e3
- PubMed: 35660158
- DOI: https://doi.org/10.1016/j.str.2022.05.009
- Primary Citation Related Structures:
7QFN, 7QFO, 7ZBG - PubMed Abstract:
Human topoisomerase II beta (TOP2B) modulates DNA topology using energy from ATP hydrolysis. To investigate the conformational changes that occur during ATP hydrolysis, we determined the X-ray crystallographic structures of the human TOP2B ATPase domain bound to AMPPNP or ADP at 1.9 Å and 2.6 Å resolution, respectively. The GHKL domains of both structures are similar, whereas the QTK loop within the transducer domain can move for product release. As TOP2B is the clinical target of bisdioxopiperazines, we also determined the structure of a TOP2B:ADP:ICRF193 complex to 2.3 Å resolution and identified key drug-binding residues. Biochemical characterization revealed the N-terminal strap reduces the rate of ATP hydrolysis. Mutagenesis demonstrated residue E103 as essential for ATP hydrolysis in TOP2B. Our data provide fundamental insights into the tertiary structure of the human TOP2B ATPase domain and a potential regulatory mechanism for ATP hydrolysis.
- Biosciences Institute, Newcastle University, Newcastle upon Tyne NE2 4HH, UK.
Organizational Affiliation: