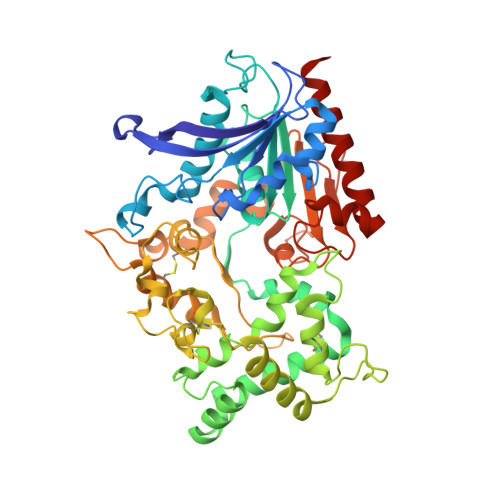
Crystal structure and substrate recognition mechanism of the prolyl endoprotease PEP from Aspergillus niger.
Miyazono, K.I., Kubota, K., Takahashi, K., Tanokura, M.(2022) Biochem Biophys Res Commun 591: 76-81
- PubMed: 34999257
- DOI: https://doi.org/10.1016/j.bbrc.2021.12.114
- Primary Citation Related Structures:
7WAB - PubMed Abstract:
Proteases are enzymes that are not only essential for life but also industrially important. Understanding the substrate recognition mechanisms of proteases is important to enhance the use of proteases. The fungus Aspergillus produces a wide variety of proteases, including PEP, which is a prolyl endoprotease from A. niger. Although PEP exhibits amino acid sequence similarity to the serine peptidase family S28 proteins (PRCP and DPP7) that recognize Pro-X bonds in the terminal regions of peptides, PEP recognizes Pro-X bonds not only in peptides but also in proteins. To reveal the structural basis of the prolyl endoprotease activity of PEP, we determined the structure of PEP by X-ray crystallography at a resolution of 1.75 Å. The PEP structure shows that PEP has a wide-open catalytic pocket compared to its homologs. The characteristic catalytic pocket structure of PEP is predicted to be important for the recognition of protein substrates.
- Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi Bunkyo-ku, Tokyo, 113-8657, Japan.
Organizational Affiliation: