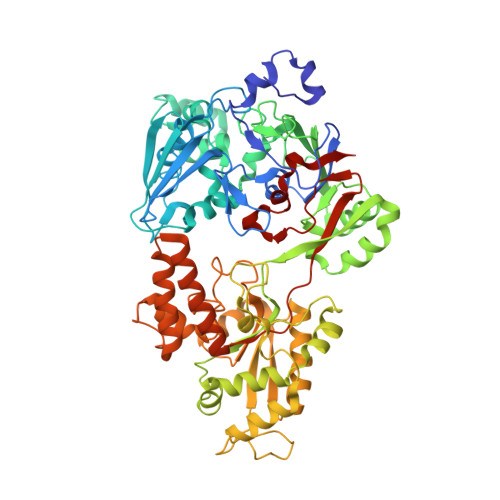
Substrate size-dependent conformational changes of bacterial pectin-binding protein crucial for chemotaxis and assimilation.
Anamizu, K., Takase, R., Hio, M., Watanabe, D., Mikami, B., Hashimoto, W.(2022) Sci Rep 12: 12653-12653
- PubMed: 35879323 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-022-16540-5
- Primary Citation Related Structures:
7VEQ, 7VER, 7VET, 7VEU, 7VEV, 7VEW - PubMed Abstract:
Gram-negative Sphingomonas sp. strain A1 exhibits positive chemotaxis toward acidic polysaccharide pectin. SPH1118 has been identified as a pectin-binding protein involved in both pectin chemotaxis and assimilation. Here we show tertiary structures of SPH1118 with six different conformations as determined by X-ray crystallography. SPH1118 consisted of two domains with a large cleft between the domains and substrates bound to positively charged and aromatic residues in the cleft through hydrogen bond and stacking interactions. Substrate-free SPH1118 adopted three different conformations in the open form. On the other hand, the two domains were closed in substrate-bound form and the domain closure ratio was changed in response to the substrate size, suggesting that the conformational change upon binding to the substrate triggered the expression of pectin chemotaxis and assimilation. This study first clarified that the solute-binding protein with dual functions recognized the substrate through flexible conformational changes in response to the substrate size.
- Laboratory of Basic and Applied Molecular Biotechnology, Division of Food Science and Biotechnology, Graduate School of Agriculture, Kyoto University, Uji, Kyoto, 611-0011, Japan.
Organizational Affiliation: