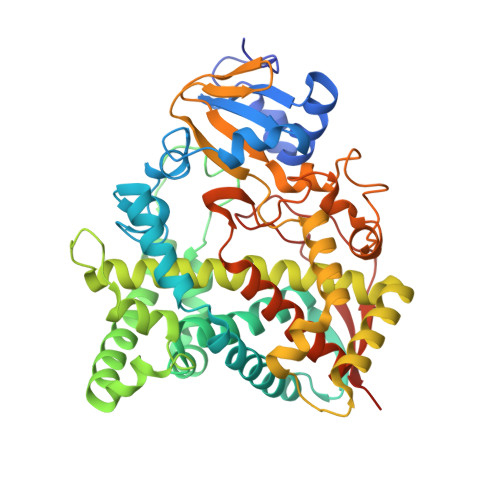
Relaxed Substrate Requirements of Sterol 14 alpha-Demethylase from Naegleria fowleri Are Accompanied by Resistance to Inhibition.
Hargrove, T.Y., Wawrzak, Z., Rachakonda, G., Nes, W.D., Villalta, F., Guengerich, F.P., Lepesheva, G.I.(2021) J Med Chem 64: 17511-17522
- PubMed: 34842434 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01710
- Primary Citation Related Structures:
7RTQ - PubMed Abstract:
Naegleria fowleri is the protozoan pathogen that causes primary amoebic meningoencephalitis (PAM), with the death rate exceeding 97%. The amoeba makes sterols and can be targeted by sterol biosynthesis inhibitors. Here, we characterized N. fowleri sterol 14-demethylase, including catalytic properties and inhibition by clinical antifungal drugs and experimental substituted azoles with favorable pharmacokinetics and low toxicity. None of them inhibited the enzyme stoichiometrically. The highest potencies were displayed by posaconazole (IC 50 = 0.69 μM) and two of our compounds (IC 50 = 1.3 and 0.35 μM). Because both these compounds penetrate the brain with concentrations reaching minimal inhibitory concentration (MIC) values in an N. fowleri cellular assay, we report them as potential drug candidates for PAM. The 2.1 Å crystal structure, in complex with the strongest inhibitor, provides an explanation connecting the enzyme weaker substrate specificity with lower sensitivity to inhibition. It also provides insight into the enzyme/ligand molecular recognition process and suggests directions for the design of more potent inhibitors.
- Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, Tennessee 37232, United States.
Organizational Affiliation: