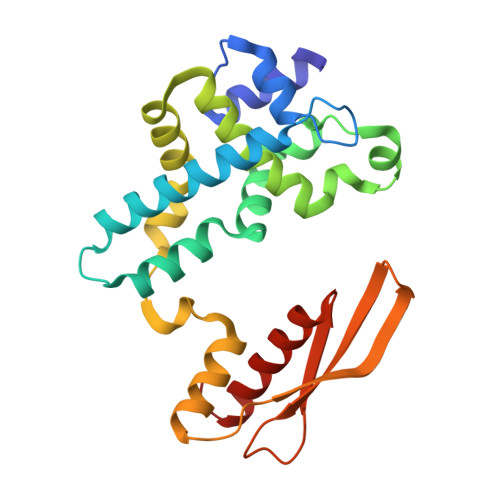
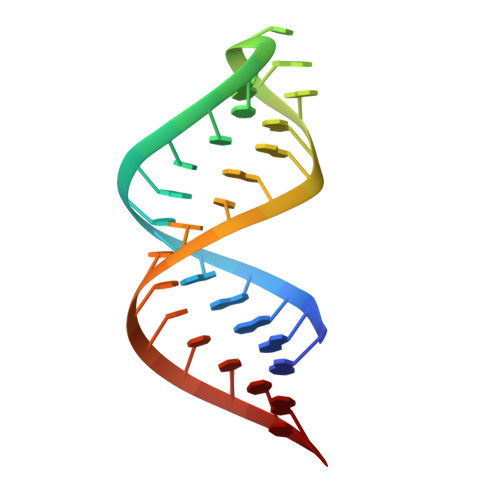
Structural basis for Dicer-like function of an engineered RNase III variant and insights into the reaction trajectory of two-Mg 2+ -ion catalysis.
Dharavath, S., Shaw, G.X., Ji, X.(2022) RNA Biol 19: 908-915
- PubMed: 35829618 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/15476286.2022.2099650
- Primary Citation Related Structures:
7R97 - PubMed Abstract:
The RNase III family of dsRNA-specific endonucleases is exemplified by prokaryotic RNase III and eukaryotic Rnt1p, Drosha, and Dicer. Structures of Aquifex aeolicus RNase III (AaRNase III) and Saccharomyces cerevisiae Rnt1p (ScRnt1p) show that both enzymes recognize substrates in a sequence-specific manner and propel RNA hydrolysis by two-Mg 2+ -ion catalysis. Previously, we created an Escherichia coli RNase III variant (EcEEQ) by eliminating the sequence specificity via protein engineering and called it bacterial Dicer for the fact that it produces heterogeneous small interfering RNA cocktails. Here, we present a 1.8-Å crystal structure of a postcleavage complex of EcEEQ, representing a reaction state immediately after the cleavage of scissile bond. The structure not only establishes the structure-and-function relationship of EcEEQ, but also reveals the functional role of a third Mg 2+ ion that is involved in RNA hydrolysis by bacterial RNase III. In contrast, the cleavage site assembly of ScRnt1p does not contain a third Mg 2+ ion. Instead, it involves two more amino acid side chains conserved among eukaryotic RNase IIIs. We conclude that the EcEEQ structure (this work) represents the cleavage assembly of prokaryotic RNase IIIs and the ScRnt1p structure (PDB: 4OOG), also determined at the postcleavage state, represents the cleavage assembly of eukaryotic RNase IIIs. Together, these two structures provide insights into the reaction trajectory of two-Mg 2+ -ion catalysis by prokaryotic and eukaryotic RNase III enzymes.
- Center for Structural Biology, National Cancer Institute, Frederick, MD, USA.
Organizational Affiliation: