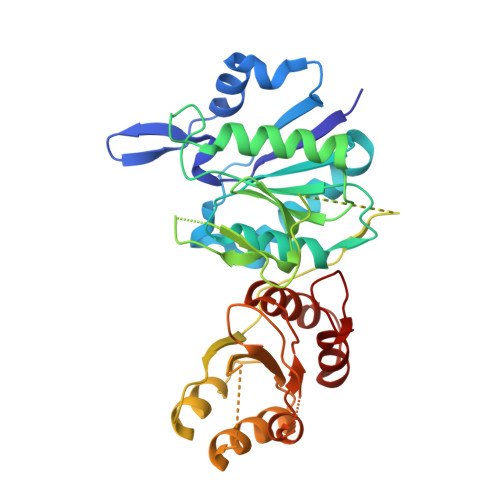
The dimeric form of bacterial l-asparaginase YpAI is fully active.
Strzelczyk, P., Zhang, D., Alexandratos, J., Piszczek, G., Wlodawer, A., Lubkowski, J.(2023) FEBS J 290: 780-795
- PubMed: 36152020 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/febs.16635
- Primary Citation Related Structures:
7R69, 7R6A, 7R6B - PubMed Abstract:
l-asparaginases from mesophilic bacteria (ASNases), including two enzymes very successfully used in the treatment of leukaemia, have been consistently described as homotetramers. On the contrary, structural studies show that homodimers of these enzymes should be sufficient to carry out the catalytic reaction. In this report, we investigated whether the type I Yersinia pestis asparaginase (YpAI) is active in a dimeric form or whether the tetrameric quaternary structure is critical for its activity. Using multiple biophysical techniques that investigate enzymatic properties and quaternary structure at either high or low protein concentration, we found that dimeric YpAI is fully active, suggesting that the tetrameric form of this subfamily of enzymes does not bear significant enzymatic relevance. In this process, we extensively characterized YpAI, showing that it is a cooperative enzyme, although the mechanism of allostery is still not definitely established. We showed that, like most type I ASNases, the substrate affinity of YpAI is low and this enzyme is very similar in terms of both the structure and enzymatic properties to homologous type I ASNase from Escherichia coli (EcAI). We extended these studies to more medically relevant type II ASNases, used as anti-leukaemia drugs. We confirmed that type II ASNases are not allosteric, and that they might also be functional in a dimeric form. However, the determination of the accurate tetramer⇆dimer dissociation constants of these enzymes that most likely lie in the picomolar range is not possible with currently available biophysical techniques.
- Center for Structural Biology, Center for Cancer Research, National Cancer Institute, Frederick, MD, USA.
Organizational Affiliation: