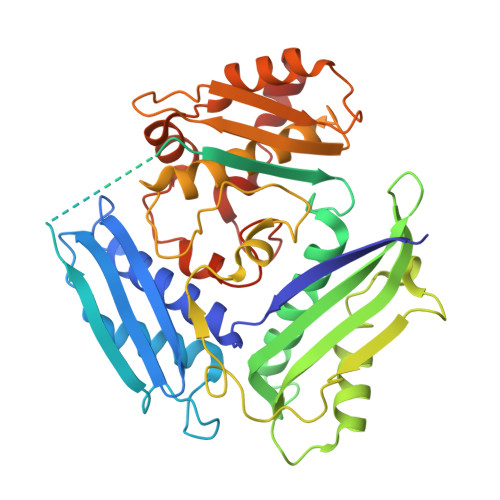
Evolution of homo-oligomerization of methionine S-adenosyltransferases is replete with structure-function constrains.
Kleiner, D., Shapiro Tuchman, Z., Shmulevich, F., Shahar, A., Zarivach, R., Kosloff, M., Bershtein, S.(2022) Protein Sci 31: e4352-e4352
- PubMed: 35762725
- DOI: https://doi.org/10.1002/pro.4352
- Primary Citation of Related Structures:
7R2W, 7R3B - PubMed Abstract:
Homomers are prevalent in bacterial proteomes, particularly among core metabolic enzymes. Homomerization is often key to function and regulation, and interfaces that facilitate the formation of homomeric enzymes are subject to intense evolutionary change. However, our understanding of the molecular mechanisms that drive evolutionary variation in homomeric complexes is still lacking. How is the diversification of protein interfaces linked to variation in functional regulation and structural integrity of homomeric complexes? To address this question, we studied quaternary structure evolution of bacterial methionine S-adenosyltransferases (MATs)-dihedral homotetramers formed along a large and conserved dimeric interface harboring two active sites, and a small, recently evolved, interdimeric interface. Here, we show that diversity in the physicochemical properties of small interfaces is directly linked to variability in the kinetic stability of MAT quaternary complexes and in modes of their functional regulation. Specifically, hydrophobic interactions within the small interface of Escherichia coli MAT render the functional homotetramer kinetically stable yet impose severe aggregation constraints on complex assembly. These constraints are alleviated by electrostatic interactions that accelerate dimer-dimer assembly. In contrast, Neisseria gonorrhoeae MAT adopts a nonfunctional dimeric state due to the low hydrophobicity of its small interface and the high flexibility of its active site loops, which perturbs small interface integrity. Remarkably, in the presence of methionine and ATP, N. gonorrhoeae MAT undergoes substrate-induced assembly into a functional tetrameric state. We suggest that evolution acts on the interdimeric interfaces of MATs to tailor the regulation of their activity and stability to unique organismal needs.
- Department of Life Sciences, Ben-Gurion University of the Negev, Beer-Sheva, Israel.
Organizational Affiliation: